fitcauto
Automatically select classification model with optimized hyperparameters
Syntax
Description
Given predictor and response data, fitcauto automatically
tries a selection of classification model types with different hyperparameter values. By
default, the function uses Bayesian optimization to select models and their hyperparameter
values, and computes the cross-validation classification error for each model. After the
optimization is complete, fitcauto returns the model, trained on the
entire data set, that is expected to best classify new data. You can use the
predict and loss object functions of the returned
model to classify new data and compute the test set classification error,
respectively.
Use fitcauto when you are uncertain which classifier types best suit
your data. For information on alternative methods for tuning hyperparameters of classification
models, see Alternative Functionality.
If your data contains over 10,000 observations, consider using an asynchronous successive
halving algorithm (ASHA) instead of Bayesian optimization when you run
fitcauto. ASHA optimization often finds good solutions faster than
Bayesian optimization for data sets with many observations.
Mdl = fitcauto(Tbl,ResponseVarName)Mdl with tuned hyperparameters. The
table Tbl contains the predictor variables and the response variable,
where ResponseVarName is the name of the response variable.
Mdl = fitcauto(___,Name,Value)HyperparameterOptimizationOptions name-value argument to specify
whether to use Bayesian optimization (default) or an asynchronous successive halving
algorithm (ASHA). To use ASHA optimization, specify
"HyperparameterOptimizationOptions",struct("Optimizer","asha"). You
can include additional fields in the structure to control other aspects of the
optimization.
[
also returns Mdl,OptimizationResults] = fitcauto(___)OptimizationResults, which contains the results of the
model selection and hyperparameter tuning process.
Examples
Use fitcauto to automatically select a classification model with optimized hyperparameters, given predictor and response data stored in a table.
Load Data
Load the carbig data set, which contains measurements of cars made in the 1970s and early 1980s.
load carbigCategorize the cars based on whether they were made in the USA.
Origin = categorical(cellstr(Origin)); Origin = mergecats(Origin,["France","Japan","Germany", ... "Sweden","Italy","England"],"NotUSA");
Create a table containing the predictor variables Acceleration, Displacement, and so on, as well as the response variable Origin.
cars = table(Acceleration,Displacement,Horsepower, ...
Model_Year,MPG,Weight,Origin);Partition Data
Partition the data into training and test sets. Use approximately 80% of the observations for the model selection and hyperparameter tuning process, and 20% of the observations to test the performance of the final model returned by fitcauto. Use cvpartition to partition the data.
rng("default") % For reproducibility of the data partition c = cvpartition(Origin,"Holdout",0.2); trainingIdx = training(c); % Training set indices carsTrain = cars(trainingIdx,:); testIdx = test(c); % Test set indices carsTest = cars(testIdx,:);
Run fitcauto
Pass the training data to fitcauto. By default, fitcauto determines appropriate model types to try, uses Bayesian optimization to find good hyperparameter values, and returns a trained model Mdl with the best expected performance. Additionally, fitcauto provides a plot of the optimization and an iterative display of the optimization results. For more information on how to interpret these results, see Verbose Display.
Expect this process to take some time. To speed up the optimization process, consider specifying to run the optimization in parallel, if you have a Parallel Computing Toolbox™ license. To do so, pass "HyperparameterOptimizationOptions",struct("UseParallel",true) to fitcauto as a name-value argument.
Mdl = fitcauto(carsTrain,"Origin");Learner types to explore: ensemble, knn, nb, net, svm, tree Total iterations (MaxObjectiveEvaluations): 180 Total time (MaxTime): Inf |=============================================================================================================================================| | Iter | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | result | loss | & validation (sec)| validation loss | validation loss | | | |=============================================================================================================================================| | 1 | Best | 0.10769 | 9.8867 | 0.10769 | 0.10769 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 291 | | | | | | | | | MinLeafSize: 4 | | 2 | Accept | 0.13231 | 8.4208 | 0.10769 | 0.12098 | ensemble | Method: Bag | | | | | | | | | NumLearningCycles: 233 | | | | | | | | | MinLeafSize: 1 | | 3 | Best | 0.10154 | 0.46968 | 0.10154 | 0.10154 | tree | MinLeafSize: 5 | | 4 | Accept | 0.37231 | 0.77391 | 0.10154 | 0.10154 | svm | BoxConstraint: 0.22029 | | | | | | | | | KernelScale: 0.4534 | | 5 | Accept | 0.14769 | 0.12722 | 0.10154 | 0.12098 | tree | MinLeafSize: 2 | | 6 | Accept | 0.24308 | 0.54475 | 0.10154 | 0.12098 | knn | NumNeighbors: 155 | | 7 | Accept | 0.37231 | 0.17065 | 0.10154 | 0.12098 | svm | BoxConstraint: 22.3 | | | | | | | | | KernelScale: 0.018132 | | 8 | Accept | 0.19385 | 0.098037 | 0.10154 | 0.12098 | knn | NumNeighbors: 75 | | 9 | Accept | 0.15692 | 8.2932 | 0.10154 | 0.12098 | net | Activations: tanh | | | | | | | | | Standardize: true | | | | | | | | | Lambda: 0.0010226 | | | | | | | | | LayerSizes: 224 | | 10 | Accept | 0.37231 | 0.20214 | 0.10154 | 0.12098 | net | Activations: tanh | | | | | | | | | Standardize: false | | | | | | | | | Lambda: 302.8 | | | | | | | | | LayerSizes: [ 78 5 ] | | 11 | Accept | 0.22769 | 1.2781 | 0.10154 | 0.12098 | nb | DistributionNames: kernel | | | | | | | | | Width: 214.96 | | | | | | | | | Standardize: false | | 12 | Accept | 0.18154 | 5.8459 | 0.10154 | 0.12323 | ensemble | Method: Bag | | | | | | | | | NumLearningCycles: 232 | | | | | | | | | MinLeafSize: 105 | | 13 | Accept | 0.19692 | 0.090905 | 0.10154 | 0.12323 | knn | NumNeighbors: 13 | | 14 | Accept | 0.20308 | 0.10957 | 0.10154 | 0.12323 | svm | BoxConstraint: 9.1458 | | | | | | | | | KernelScale: 18.65 | | 15 | Accept | 0.14154 | 7.7326 | 0.10154 | 0.12323 | ensemble | Method: Bag | | | | | | | | | NumLearningCycles: 278 | | | | | | | | | MinLeafSize: 2 | | 16 | Accept | 0.11692 | 4.5938 | 0.10154 | 0.12323 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 214 | | | | | | | | | MinLeafSize: 1 | | 17 | Accept | 0.18462 | 0.088159 | 0.10154 | 0.12323 | knn | NumNeighbors: 10 | | 18 | Accept | 0.12308 | 0.077702 | 0.10154 | 0.12484 | tree | MinLeafSize: 10 | | 19 | Accept | 0.26462 | 3.68 | 0.10154 | 0.12484 | net | Activations: tanh | | | | | | | | | Standardize: false | | | | | | | | | Lambda: 0.44352 | | | | | | | | | LayerSizes: [ 48 40 ] | | 20 | Accept | 0.10769 | 4.1582 | 0.10154 | 0.12484 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 206 | | | | | | | | | MinLeafSize: 3 | |=============================================================================================================================================| | Iter | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | result | loss | & validation (sec)| validation loss | validation loss | | | |=============================================================================================================================================| | 21 | Accept | 0.12 | 0.092835 | 0.10154 | 0.12349 | tree | MinLeafSize: 13 | | 22 | Accept | 0.19385 | 0.08931 | 0.10154 | 0.12349 | knn | NumNeighbors: 57 | | 23 | Accept | 0.37231 | 0.29219 | 0.10154 | 0.12349 | net | Activations: relu | | | | | | | | | Standardize: true | | | | | | | | | Lambda: 1.6049 | | | | | | | | | LayerSizes: [ 236 2 197 ] | | 24 | Accept | 0.13538 | 7.2642 | 0.10154 | 0.12349 | ensemble | Method: Bag | | | | | | | | | NumLearningCycles: 272 | | | | | | | | | MinLeafSize: 4 | | 25 | Accept | 0.21231 | 0.12119 | 0.10154 | 0.12349 | svm | BoxConstraint: 3.2832 | | | | | | | | | KernelScale: 19.127 | | 26 | Accept | 0.23692 | 0.088582 | 0.10154 | 0.12349 | knn | NumNeighbors: 154 | | 27 | Accept | 0.18154 | 0.071505 | 0.10154 | 0.12413 | tree | MinLeafSize: 30 | | 28 | Accept | 0.22769 | 0.14963 | 0.10154 | 0.12413 | nb | DistributionNames: normal | | | | | | | | | Width: NaN | | | | | | | | | Standardize: - | | 29 | Accept | 0.37231 | 0.31305 | 0.10154 | 0.12413 | net | Activations: tanh | | | | | | | | | Standardize: false | | | | | | | | | Lambda: 9.7449 | | | | | | | | | LayerSizes: [ 123 213 22 ] | | 30 | Accept | 0.22769 | 0.061021 | 0.10154 | 0.12413 | nb | DistributionNames: normal | | | | | | | | | Width: NaN | | | | | | | | | Standardize: - | | 31 | Accept | 0.20615 | 7.6661 | 0.10154 | 0.12413 | net | Activations: sigmoid | | | | | | | | | Standardize: false | | | | | | | | | Lambda: 9.0476e-06 | | | | | | | | | LayerSizes: [ 1 286 ] | | 32 | Accept | 0.37231 | 0.12123 | 0.10154 | 0.12413 | net | Activations: none | | | | | | | | | Standardize: true | | | | | | | | | Lambda: 169.93 | | | | | | | | | LayerSizes: [ 8 43 ] | | 33 | Accept | 0.16923 | 0.053687 | 0.10154 | 0.12413 | tree | MinLeafSize: 50 | | 34 | Accept | 0.12923 | 0.052139 | 0.10154 | 0.12413 | tree | MinLeafSize: 3 | | 35 | Accept | 0.37538 | 0.16254 | 0.10154 | 0.12413 | svm | BoxConstraint: 31.988 | | | | | | | | | KernelScale: 0.068262 | | 36 | Best | 0.086154 | 5.1664 | 0.086154 | 0.11432 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 259 | | | | | | | | | MinLeafSize: 46 | | 37 | Accept | 0.20308 | 0.093464 | 0.086154 | 0.11432 | knn | NumNeighbors: 7 | | 38 | Accept | 0.19692 | 0.062179 | 0.086154 | 0.11432 | knn | NumNeighbors: 9 | | 39 | Accept | 0.21846 | 0.20379 | 0.086154 | 0.11432 | nb | DistributionNames: kernel | | | | | | | | | Width: 11.43 | | | | | | | | | Standardize: false | | 40 | Accept | 0.22769 | 0.077831 | 0.086154 | 0.11432 | nb | DistributionNames: normal | | | | | | | | | Width: NaN | | | | | | | | | Standardize: - | |=============================================================================================================================================| | Iter | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | result | loss | & validation (sec)| validation loss | validation loss | | | |=============================================================================================================================================| | 41 | Accept | 0.24615 | 0.2016 | 0.086154 | 0.11432 | nb | DistributionNames: kernel | | | | | | | | | Width: 0.95982 | | | | | | | | | Standardize: false | | 42 | Accept | 0.22769 | 0.054205 | 0.086154 | 0.11432 | nb | DistributionNames: normal | | | | | | | | | Width: NaN | | | | | | | | | Standardize: - | | 43 | Accept | 0.16308 | 1.8399 | 0.086154 | 0.11432 | net | Activations: tanh | | | | | | | | | Standardize: true | | | | | | | | | Lambda: 8.2888e-08 | | | | | | | | | LayerSizes: 20 | | 44 | Accept | 0.37231 | 0.10773 | 0.086154 | 0.11432 | svm | BoxConstraint: 0.5593 | | | | | | | | | KernelScale: 0.003667 | | 45 | Accept | 0.11385 | 0.075454 | 0.086154 | 0.11432 | tree | MinLeafSize: 11 | | 46 | Accept | 0.20308 | 0.074108 | 0.086154 | 0.11432 | knn | NumNeighbors: 7 | | 47 | Accept | 0.22769 | 0.061325 | 0.086154 | 0.11432 | nb | DistributionNames: normal | | | | | | | | | Width: NaN | | | | | | | | | Standardize: - | | 48 | Accept | 0.25231 | 5.6001 | 0.086154 | 0.11432 | net | Activations: sigmoid | | | | | | | | | Standardize: false | | | | | | | | | Lambda: 4.2329e-05 | | | | | | | | | LayerSizes: [ 16 21 86 ] | | 49 | Accept | 0.22769 | 0.051748 | 0.086154 | 0.11432 | nb | DistributionNames: normal | | | | | | | | | Width: NaN | | | | | | | | | Standardize: - | | 50 | Accept | 0.37231 | 0.11468 | 0.086154 | 0.11432 | svm | BoxConstraint: 47.547 | | | | | | | | | KernelScale: 0.038904 | | 51 | Accept | 0.16923 | 0.074099 | 0.086154 | 0.11432 | tree | MinLeafSize: 76 | | 52 | Accept | 0.15692 | 1.6882 | 0.086154 | 0.11432 | net | Activations: tanh | | | | | | | | | Standardize: false | | | | | | | | | Lambda: 0.24612 | | | | | | | | | LayerSizes: 6 | | 53 | Accept | 0.10462 | 4.0245 | 0.086154 | 0.10962 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 211 | | | | | | | | | MinLeafSize: 6 | | 54 | Accept | 0.11385 | 0.092927 | 0.086154 | 0.10962 | tree | MinLeafSize: 11 | | 55 | Accept | 0.086154 | 5.1988 | 0.086154 | 0.098594 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 286 | | | | | | | | | MinLeafSize: 47 | | 56 | Accept | 0.19692 | 0.075963 | 0.086154 | 0.098594 | knn | NumNeighbors: 28 | | 57 | Accept | 0.12 | 0.070258 | 0.086154 | 0.098594 | tree | MinLeafSize: 4 | | 58 | Accept | 0.2 | 0.068373 | 0.086154 | 0.098594 | knn | NumNeighbors: 3 | | 59 | Accept | 0.37231 | 0.15296 | 0.086154 | 0.098594 | net | Activations: tanh | | | | | | | | | Standardize: false | | | | | | | | | Lambda: 79.211 | | | | | | | | | LayerSizes: 1 | | 60 | Accept | 0.19692 | 0.061209 | 0.086154 | 0.098594 | knn | NumNeighbors: 13 | |=============================================================================================================================================| | Iter | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | result | loss | & validation (sec)| validation loss | validation loss | | | |=============================================================================================================================================| | 61 | Accept | 0.37231 | 5.3523 | 0.086154 | 0.10784 | ensemble | Method: Bag | | | | | | | | | NumLearningCycles: 234 | | | | | | | | | MinLeafSize: 155 | | 62 | Accept | 0.21846 | 0.076683 | 0.086154 | 0.10784 | knn | NumNeighbors: 2 | | 63 | Accept | 0.32308 | 1.0531 | 0.086154 | 0.10784 | net | Activations: none | | | | | | | | | Standardize: false | | | | | | | | | Lambda: 0.00049454 | | | | | | | | | LayerSizes: [ 3 145 ] | | 64 | Accept | 0.11077 | 5.6836 | 0.086154 | 0.10698 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 300 | | | | | | | | | MinLeafSize: 1 | | 65 | Accept | 0.19692 | 2.5205 | 0.086154 | 0.10698 | net | Activations: tanh | | | | | | | | | Standardize: true | | | | | | | | | Lambda: 1.1014e-05 | | | | | | | | | LayerSizes: [ 26 2 ] | | 66 | Accept | 0.12 | 0.072271 | 0.086154 | 0.10698 | tree | MinLeafSize: 4 | | 67 | Accept | 0.21846 | 0.095838 | 0.086154 | 0.10698 | knn | NumNeighbors: 2 | | 68 | Accept | 0.37231 | 0.091219 | 0.086154 | 0.10698 | svm | BoxConstraint: 0.0061987 | | | | | | | | | KernelScale: 31.312 | | 69 | Accept | 0.37231 | 0.064588 | 0.086154 | 0.10698 | svm | BoxConstraint: 0.070644 | | | | | | | | | KernelScale: 0.22263 | | 70 | Accept | 0.37538 | 0.082788 | 0.086154 | 0.10698 | svm | BoxConstraint: 9.2391 | | | | | | | | | KernelScale: 0.062892 | | 71 | Accept | 0.15692 | 0.061431 | 0.086154 | 0.10698 | svm | BoxConstraint: 32.735 | | | | | | | | | KernelScale: 1.3401 | | 72 | Accept | 0.31077 | 0.05396 | 0.086154 | 0.10698 | svm | BoxConstraint: 0.0407 | | | | | | | | | KernelScale: 3.4118 | | 73 | Accept | 0.37231 | 0.053805 | 0.086154 | 0.10698 | svm | BoxConstraint: 0.0019745 | | | | | | | | | KernelScale: 62.689 | | 74 | Accept | 0.13538 | 5.1971 | 0.086154 | 0.10699 | ensemble | Method: Bag | | | | | | | | | NumLearningCycles: 205 | | | | | | | | | MinLeafSize: 3 | | 75 | Accept | 0.22769 | 0.068918 | 0.086154 | 0.10699 | nb | DistributionNames: normal | | | | | | | | | Width: NaN | | | | | | | | | Standardize: - | | 76 | Accept | 0.37231 | 0.06363 | 0.086154 | 0.10699 | svm | BoxConstraint: 0.42987 | | | | | | | | | KernelScale: 234.78 | | 77 | Accept | 0.25846 | 0.2272 | 0.086154 | 0.10699 | nb | DistributionNames: kernel | | | | | | | | | Width: 2.2007 | | | | | | | | | Standardize: false | | 78 | Accept | 0.13846 | 5.0838 | 0.086154 | 0.10344 | ensemble | Method: Bag | | | | | | | | | NumLearningCycles: 207 | | | | | | | | | MinLeafSize: 4 | | 79 | Accept | 0.14462 | 0.071561 | 0.086154 | 0.10344 | tree | MinLeafSize: 21 | | 80 | Accept | 0.37231 | 0.097395 | 0.086154 | 0.10344 | svm | BoxConstraint: 176.96 | | | | | | | | | KernelScale: 0.022732 | |=============================================================================================================================================| | Iter | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | result | loss | & validation (sec)| validation loss | validation loss | | | |=============================================================================================================================================| | 81 | Accept | 0.19692 | 0.07841 | 0.086154 | 0.10344 | knn | NumNeighbors: 9 | | 82 | Accept | 0.26154 | 0.17034 | 0.086154 | 0.10344 | nb | DistributionNames: kernel | | | | | | | | | Width: 2.808 | | | | | | | | | Standardize: false | | 83 | Accept | 0.37231 | 0.18007 | 0.086154 | 0.10344 | nb | DistributionNames: kernel | | | | | | | | | Width: 615.95 | | | | | | | | | Standardize: true | | 84 | Accept | 0.26154 | 1.8058 | 0.086154 | 0.10344 | net | Activations: relu | | | | | | | | | Standardize: false | | | | | | | | | Lambda: 8.2545e-08 | | | | | | | | | LayerSizes: [ 57 2 7 ] | | 85 | Accept | 0.22769 | 0.063696 | 0.086154 | 0.10344 | nb | DistributionNames: normal | | | | | | | | | Width: NaN | | | | | | | | | Standardize: - | | 86 | Accept | 0.14769 | 6.2743 | 0.086154 | 0.10769 | ensemble | Method: Bag | | | | | | | | | NumLearningCycles: 252 | | | | | | | | | MinLeafSize: 32 | | 87 | Accept | 0.17231 | 0.06519 | 0.086154 | 0.10769 | tree | MinLeafSize: 126 | | 88 | Accept | 0.16923 | 0.043935 | 0.086154 | 0.10769 | tree | MinLeafSize: 93 | | 89 | Accept | 0.22769 | 0.18302 | 0.086154 | 0.10769 | nb | DistributionNames: kernel | | | | | | | | | Width: 0.43629 | | | | | | | | | Standardize: false | | 90 | Accept | 0.26462 | 3.9886 | 0.086154 | 0.10769 | net | Activations: sigmoid | | | | | | | | | Standardize: false | | | | | | | | | Lambda: 7.2789e-07 | | | | | | | | | LayerSizes: 98 | | 91 | Accept | 0.098462 | 4.8539 | 0.086154 | 0.10445 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 259 | | | | | | | | | MinLeafSize: 39 | | 92 | Accept | 0.11077 | 4.9109 | 0.086154 | 0.099362 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 273 | | | | | | | | | MinLeafSize: 70 | | 93 | Accept | 0.13846 | 5.1708 | 0.086154 | 0.10034 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 281 | | | | | | | | | MinLeafSize: 84 | | 94 | Accept | 0.12615 | 4.9036 | 0.086154 | 0.099454 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 244 | | | | | | | | | MinLeafSize: 76 | | 95 | Accept | 0.11077 | 5.5508 | 0.086154 | 0.097618 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 285 | | | | | | | | | MinLeafSize: 71 | | 96 | Accept | 0.14154 | 4.4952 | 0.086154 | 0.095919 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 245 | | | | | | | | | MinLeafSize: 57 | | 97 | Accept | 0.095385 | 4.815 | 0.086154 | 0.094528 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 260 | | | | | | | | | MinLeafSize: 51 | | 98 | Accept | 0.12308 | 4.6592 | 0.086154 | 0.093172 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 260 | | | | | | | | | MinLeafSize: 77 | | 99 | Accept | 0.37231 | 4.8941 | 0.086154 | 0.094437 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 268 | | | | | | | | | MinLeafSize: 149 | | 100 | Accept | 0.13538 | 5.2577 | 0.086154 | 0.096586 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 295 | | | | | | | | | MinLeafSize: 92 | |=============================================================================================================================================| | Iter | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | result | loss | & validation (sec)| validation loss | validation loss | | | |=============================================================================================================================================| | 101 | Accept | 0.092308 | 4.4168 | 0.086154 | 0.092045 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 247 | | | | | | | | | MinLeafSize: 43 | | 102 | Accept | 0.13846 | 4.8009 | 0.086154 | 0.093381 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 265 | | | | | | | | | MinLeafSize: 90 | | 103 | Accept | 0.089231 | 4.393 | 0.086154 | 0.091453 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 245 | | | | | | | | | MinLeafSize: 48 | | 104 | Accept | 0.098462 | 4.7966 | 0.086154 | 0.091605 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 265 | | | | | | | | | MinLeafSize: 52 | | 105 | Accept | 0.086154 | 4.9362 | 0.086154 | 0.090054 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 276 | | | | | | | | | MinLeafSize: 47 | | 106 | Accept | 0.089231 | 4.1084 | 0.086154 | 0.088948 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 224 | | | | | | | | | MinLeafSize: 45 | | 107 | Accept | 0.13538 | 5.0023 | 0.086154 | 0.089673 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 279 | | | | | | | | | MinLeafSize: 97 | | 108 | Accept | 0.13538 | 5.0726 | 0.086154 | 0.089434 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 284 | | | | | | | | | MinLeafSize: 99 | | 109 | Accept | 0.14154 | 3.9115 | 0.086154 | 0.089922 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 214 | | | | | | | | | MinLeafSize: 99 | | 110 | Accept | 0.089231 | 5.324 | 0.086154 | 0.089121 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 296 | | | | | | | | | MinLeafSize: 47 | | 111 | Accept | 0.18154 | 5.2688 | 0.086154 | 0.088882 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 293 | | | | | | | | | MinLeafSize: 124 | | 112 | Accept | 0.13231 | 4.3796 | 0.086154 | 0.088506 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 247 | | | | | | | | | MinLeafSize: 54 | | 113 | Accept | 0.098462 | 3.9631 | 0.086154 | 0.087879 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 219 | | | | | | | | | MinLeafSize: 51 | | 114 | Accept | 0.11077 | 5.0266 | 0.086154 | 0.088112 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 278 | | | | | | | | | MinLeafSize: 28 | | 115 | Accept | 0.098462 | 5.1256 | 0.086154 | 0.089033 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 286 | | | | | | | | | MinLeafSize: 37 | | 116 | Accept | 0.089231 | 5.3373 | 0.086154 | 0.08886 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 298 | | | | | | | | | MinLeafSize: 47 | | 117 | Accept | 0.086154 | 4.9772 | 0.086154 | 0.087554 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 273 | | | | | | | | | MinLeafSize: 47 | | 118 | Accept | 0.095385 | 4.9785 | 0.086154 | 0.087693 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 276 | | | | | | | | | MinLeafSize: 46 | | 119 | Accept | 0.086154 | 5.4771 | 0.086154 | 0.087702 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 283 | | | | | | | | | MinLeafSize: 49 | | 120 | Accept | 0.092308 | 3.7618 | 0.086154 | 0.087509 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 205 | | | | | | | | | MinLeafSize: 47 | |=============================================================================================================================================| | Iter | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | result | loss | & validation (sec)| validation loss | validation loss | | | |=============================================================================================================================================| | 121 | Accept | 0.10462 | 4.2801 | 0.086154 | 0.087866 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 236 | | | | | | | | | MinLeafSize: 52 | | 122 | Accept | 0.37231 | 0.10976 | 0.086154 | 0.087866 | net | Activations: tanh | | | | | | | | | Standardize: true | | | | | | | | | Lambda: 0.72122 | | | | | | | | | LayerSizes: 13 | | 123 | Accept | 0.37231 | 0.10272 | 0.086154 | 0.087866 | net | Activations: tanh | | | | | | | | | Standardize: true | | | | | | | | | Lambda: 0.43917 | | | | | | | | | LayerSizes: 2 | | 124 | Accept | 0.2 | 0.12343 | 0.086154 | 0.087866 | net | Activations: tanh | | | | | | | | | Standardize: true | | | | | | | | | Lambda: 0.050009 | | | | | | | | | LayerSizes: 6 | | 125 | Accept | 0.29231 | 0.12056 | 0.086154 | 0.087866 | net | Activations: tanh | | | | | | | | | Standardize: true | | | | | | | | | Lambda: 0.30341 | | | | | | | | | LayerSizes: 8 | | 126 | Accept | 0.089231 | 4.654 | 0.086154 | 0.087499 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 259 | | | | | | | | | MinLeafSize: 48 | | 127 | Accept | 0.086154 | 5.012 | 0.086154 | 0.086595 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 282 | | | | | | | | | MinLeafSize: 49 | | 128 | Accept | 0.18154 | 4.9285 | 0.086154 | 0.08742 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 282 | | | | | | | | | MinLeafSize: 124 | | 129 | Accept | 0.086154 | 5.1215 | 0.086154 | 0.087394 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 277 | | | | | | | | | MinLeafSize: 49 | | 130 | Accept | 0.086154 | 4.8293 | 0.086154 | 0.086574 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 270 | | | | | | | | | MinLeafSize: 49 | | 131 | Accept | 0.13846 | 4.2622 | 0.086154 | 0.086693 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 239 | | | | | | | | | MinLeafSize: 104 | | 132 | Accept | 0.14154 | 4.4707 | 0.086154 | 0.086428 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 252 | | | | | | | | | MinLeafSize: 104 | | 133 | Accept | 0.37231 | 0.23993 | 0.086154 | 0.086428 | net | Activations: tanh | | | | | | | | | Standardize: true | | | | | | | | | Lambda: 0.15163 | | | | | | | | | LayerSizes: [ 20 93 ] | | 134 | Best | 0.083077 | 4.9394 | 0.083077 | 0.085629 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 275 | | | | | | | | | MinLeafSize: 49 | | 135 | Accept | 0.086154 | 4.7512 | 0.083077 | 0.08537 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 266 | | | | | | | | | MinLeafSize: 49 | | 136 | Accept | 0.089231 | 4.1396 | 0.083077 | 0.085748 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 228 | | | | | | | | | MinLeafSize: 49 | | 137 | Accept | 0.14154 | 4.5844 | 0.083077 | 0.08557 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 263 | | | | | | | | | MinLeafSize: 110 | | 138 | Accept | 0.089231 | 4.1878 | 0.083077 | 0.085505 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 233 | | | | | | | | | MinLeafSize: 49 | | 139 | Accept | 0.15385 | 6.4706 | 0.083077 | 0.085674 | ensemble | Method: Bag | | | | | | | | | NumLearningCycles: 273 | | | | | | | | | MinLeafSize: 49 | | 140 | Accept | 0.17538 | 6.311 | 0.083077 | 0.08568 | ensemble | Method: Bag | | | | | | | | | NumLearningCycles: 271 | | | | | | | | | MinLeafSize: 106 | |=============================================================================================================================================| | Iter | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | result | loss | & validation (sec)| validation loss | validation loss | | | |=============================================================================================================================================| | 141 | Accept | 0.083077 | 5.0683 | 0.083077 | 0.085124 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 275 | | | | | | | | | MinLeafSize: 49 | | 142 | Accept | 0.083077 | 4.9466 | 0.083077 | 0.084346 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 272 | | | | | | | | | MinLeafSize: 49 | | 143 | Accept | 0.18462 | 4.0363 | 0.083077 | 0.084296 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 227 | | | | | | | | | MinLeafSize: 115 | | 144 | Accept | 0.098462 | 4.8689 | 0.083077 | 0.084396 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 271 | | | | | | | | | MinLeafSize: 50 | | 145 | Accept | 0.17231 | 0.11083 | 0.083077 | 0.084396 | svm | BoxConstraint: 7.4019 | | | | | | | | | KernelScale: 0.98931 | | 146 | Accept | 0.21538 | 0.059402 | 0.083077 | 0.084396 | svm | BoxConstraint: 17.066 | | | | | | | | | KernelScale: 0.77684 | | 147 | Accept | 0.086154 | 4.8705 | 0.083077 | 0.084203 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 271 | | | | | | | | | MinLeafSize: 48 | | 148 | Accept | 0.086154 | 4.8211 | 0.083077 | 0.084345 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 270 | | | | | | | | | MinLeafSize: 49 | | 149 | Accept | 0.37231 | 0.063284 | 0.083077 | 0.084345 | svm | BoxConstraint: 41.79 | | | | | | | | | KernelScale: 281.24 | | 150 | Accept | 0.21846 | 0.06502 | 0.083077 | 0.084345 | svm | BoxConstraint: 59.071 | | | | | | | | | KernelScale: 0.64404 | | 151 | Accept | 0.21231 | 0.058461 | 0.083077 | 0.084345 | svm | BoxConstraint: 27.03 | | | | | | | | | KernelScale: 0.58467 | | 152 | Accept | 0.16 | 0.054338 | 0.083077 | 0.084345 | svm | BoxConstraint: 1.0872 | | | | | | | | | KernelScale: 1.1762 | | 153 | Accept | 0.17846 | 0.055672 | 0.083077 | 0.084345 | svm | BoxConstraint: 0.55409 | | | | | | | | | KernelScale: 1.0383 | | 154 | Accept | 0.37231 | 0.056181 | 0.083077 | 0.084345 | svm | BoxConstraint: 0.002375 | | | | | | | | | KernelScale: 1.1605 | | 155 | Accept | 0.16923 | 0.057591 | 0.083077 | 0.084345 | svm | BoxConstraint: 0.5264 | | | | | | | | | KernelScale: 1.7249 | | 156 | Accept | 0.16308 | 0.076268 | 0.083077 | 0.084345 | svm | BoxConstraint: 0.4426 | | | | | | | | | KernelScale: 1.3849 | | 157 | Accept | 0.16308 | 0.071124 | 0.083077 | 0.084345 | svm | BoxConstraint: 0.43171 | | | | | | | | | KernelScale: 1.2674 | | 158 | Accept | 0.17231 | 0.059062 | 0.083077 | 0.084345 | svm | BoxConstraint: 0.33464 | | | | | | | | | KernelScale: 1.3315 | | 159 | Accept | 0.19385 | 0.069434 | 0.083077 | 0.084345 | svm | BoxConstraint: 0.15356 | | | | | | | | | KernelScale: 1.9858 | | 160 | Accept | 0.20308 | 0.057155 | 0.083077 | 0.084345 | svm | BoxConstraint: 0.1477 | | | | | | | | | KernelScale: 1.5045 | |=============================================================================================================================================| | Iter | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | result | loss | & validation (sec)| validation loss | validation loss | | | |=============================================================================================================================================| | 161 | Accept | 0.23692 | 0.062405 | 0.083077 | 0.084345 | svm | BoxConstraint: 0.42828 | | | | | | | | | KernelScale: 14.853 | | 162 | Accept | 0.086154 | 5.1922 | 0.083077 | 0.084288 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 275 | | | | | | | | | MinLeafSize: 47 | | 163 | Accept | 0.083077 | 5.2334 | 0.083077 | 0.084062 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 287 | | | | | | | | | MinLeafSize: 49 | | 164 | Accept | 0.20923 | 0.066873 | 0.083077 | 0.084062 | svm | BoxConstraint: 1.6705 | | | | | | | | | KernelScale: 0.51936 | | 165 | Accept | 0.083077 | 4.8932 | 0.083077 | 0.083699 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 274 | | | | | | | | | MinLeafSize: 48 | | 166 | Accept | 0.086154 | 5.2371 | 0.083077 | 0.083963 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 296 | | | | | | | | | MinLeafSize: 49 | | 167 | Accept | 0.37231 | 0.067167 | 0.083077 | 0.083963 | svm | BoxConstraint: 0.0024433 | | | | | | | | | KernelScale: 1.378 | | 168 | Accept | 0.32923 | 0.067094 | 0.083077 | 0.083963 | svm | BoxConstraint: 0.62588 | | | | | | | | | KernelScale: 0.49029 | | 169 | Accept | 0.19077 | 0.068847 | 0.083077 | 0.083963 | svm | BoxConstraint: 0.2494 | | | | | | | | | KernelScale: 1.2705 | | 170 | Accept | 0.24 | 0.055606 | 0.083077 | 0.083963 | svm | BoxConstraint: 0.85565 | | | | | | | | | KernelScale: 0.50742 | | 171 | Accept | 0.12923 | 3.9215 | 0.083077 | 0.083927 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 218 | | | | | | | | | MinLeafSize: 53 | | 172 | Accept | 0.11077 | 5.1249 | 0.083077 | 0.083981 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 283 | | | | | | | | | MinLeafSize: 34 | | 173 | Accept | 0.37231 | 6.449 | 0.083077 | 0.083855 | ensemble | Method: Bag | | | | | | | | | NumLearningCycles: 298 | | | | | | | | | MinLeafSize: 137 | | 174 | Accept | 0.20615 | 4.7277 | 0.083077 | 0.083837 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 263 | | | | | | | | | MinLeafSize: 128 | | 175 | Best | 0.083077 | 4.9307 | 0.083077 | 0.083627 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 274 | | | | | | | | | MinLeafSize: 44 | | 176 | Accept | 0.089231 | 4.9299 | 0.083077 | 0.083892 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 274 | | | | | | | | | MinLeafSize: 45 | | 177 | Accept | 0.092308 | 4.9475 | 0.083077 | 0.083668 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 274 | | | | | | | | | MinLeafSize: 52 | | 178 | Accept | 0.086154 | 5.237 | 0.083077 | 0.08391 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 291 | | | | | | | | | MinLeafSize: 49 | | 179 | Accept | 0.083077 | 4.8488 | 0.083077 | 0.08378 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 272 | | | | | | | | | MinLeafSize: 49 | | 180 | Accept | 0.083077 | 4.9095 | 0.083077 | 0.083649 | ensemble | Method: LogitBoost | | | | | | | | | NumLearningCycles: 274 | | | | | | | | | MinLeafSize: 48 | __________________________________________________________ Optimization completed. Total iterations: 180 Total elapsed time: 624.9642 seconds Total time for training and validation: 456.1613 seconds Best observed learner is an ensemble model with: Learner: ensemble Method: LogitBoost NumLearningCycles: 274 MinLeafSize: 44 Observed validation loss: 0.083077 Time for training and validation: 4.9307 seconds Best estimated learner (returned model) is an ensemble model with: Learner: ensemble Method: LogitBoost NumLearningCycles: 275 MinLeafSize: 49 Estimated validation loss: 0.083649 Estimated time for training and validation: 5.0351 seconds Documentation for fitcauto display

The final model returned by fitcauto corresponds to the best estimated learner. Before returning the model, the function retrains it using the entire training data (carsTrain), the listed Learner (or model) type, and the displayed hyperparameter values.
Evaluate Test Set Performance
Evaluate the performance of the model on the test set.
testAccuracy = 1 - loss(Mdl,carsTest,"Origin")testAccuracy = 0.9389
confusionchart(carsTest.Origin,predict(Mdl,carsTest))
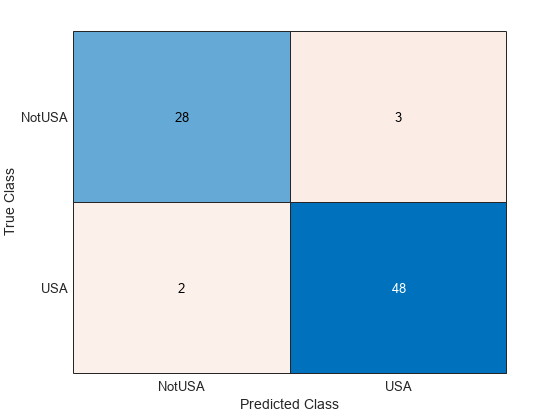
Use fitcauto to automatically select a classification model with optimized hyperparameters, given predictor and response data stored in separate variables.
Load Data
Load the humanactivity data set. This data set contains 24,075 observations of five physical human activities: Sitting (1), Standing (2), Walking (3), Running (4), and Dancing (5). Each observation has 60 features extracted from acceleration data measured by smartphone accelerometer sensors. The variable feat contains the predictor data matrix of the 60 features for the 24,075 observations, and the response variable actid contains the activity IDs for the observations as integers.
load humanactivityPartition Data
Partition the data into training and test sets. Use 90% of the observations to select a model, and 10% of the observations to validate the final model returned by fitcauto. Use cvpartition to reserve 10% of the observations for testing.
rng("default") % For reproducibility of the partition c = cvpartition(actid,"Holdout",0.10); trainingIndices = training(c); % Indices for the training set XTrain = feat(trainingIndices,:); YTrain = actid(trainingIndices); testIndices = test(c); % Indices for the test set XTest = feat(testIndices,:); YTest = actid(testIndices);
Run fitcauto
Pass the training data to fitcauto. Because the training data XTrain has more than 10,000 observations, use ASHA optimization rather than Bayesian optimization. The fitcauto function randomly selects appropriate model (or learner) types with different hyperparameter values, trains the models on a small subset of the training data, promotes the models that perform well, and retrains the promoted models on progressively larger sets of training data. The function returns the model with the best cross-validation performance, retrained on all the training data, and a table that contains the details of the optimization. Specify to run the optimization in parallel (requires Parallel Computing Toolbox™).
By default, fitcauto provides a plot of the optimization and an iterative display of the optimization results. For more information on how to interpret these results, see Verbose Display.
options = struct("Optimizer","asha","UseParallel",true); [Mdl,OptimizationResults] = fitcauto(XTrain,YTrain,"HyperparameterOptimizationOptions",options);
Starting parallel pool (parpool) using the 'Processes' profile ... 10-May-2024 09:40:17: Job Queued. Waiting for parallel pool job with ID 1 to start ... Connected to parallel pool with 6 workers. Copying objective function to workers... Done copying objective function to workers. Learner types to explore: ensemble, knn, nb, net, svm, tree Total iterations (MaxObjectiveEvaluations): 595 Total time (MaxTime): Inf |====================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Training set | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | size | | | |====================================================================================================================================================| | 1 | 6 | Best | 0.056535 | 4.0512 | 0.056535 | 271 | tree | MinLeafSize: 1 | | 2 | 6 | Accept | 0.74165 | 4.8665 | 0.056535 | 271 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 1.545 | | | | | | | | | | LayerSizes: 156 | | 3 | 6 | Accept | 0.094794 | 3.0045 | 0.056535 | 271 | knn | NumNeighbors: 27 | | 4 | 6 | Accept | 0.74165 | 8.1847 | 0.056535 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 0.0019682 | | | | | | | | | | KernelScale: 87.187 | | 5 | 4 | Accept | 0.74165 | 8.8076 | 0.056535 | 271 | knn | NumNeighbors: 1634 | | 6 | 4 | Accept | 0.74165 | 8.2922 | 0.056535 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 0.012947 | | | | | | | | | | KernelScale: 45.176 | | 7 | 4 | Accept | 0.74165 | 8.4398 | 0.056535 | 271 | knn | NumNeighbors: 375 | | 8 | 6 | Accept | 0.057873 | 4.062 | 0.056535 | 271 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.0016149 | | | | | | | | | | LayerSizes: 18 | | 9 | 6 | Best | 0.049243 | 2.4566 | 0.049243 | 1084 | tree | MinLeafSize: 1 | | 10 | 6 | Accept | 0.062719 | 3.6253 | 0.049243 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 11 | 6 | Accept | 0.74165 | 8.1485 | 0.049243 | 271 | knn | NumNeighbors: 331 | | 12 | 6 | Accept | 0.74165 | 5.4441 | 0.049243 | 271 | knn | NumNeighbors: 3719 | | 13 | 6 | Best | 0.032075 | 13.099 | 0.032075 | 1084 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.0016149 | | | | | | | | | | LayerSizes: 18 | | 14 | 6 | Accept | 0.064934 | 1.1667 | 0.032075 | 271 | knn | NumNeighbors: 2 | | 15 | 6 | Accept | 0.050858 | 1.3991 | 0.032075 | 1084 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 16 | 6 | Accept | 0.74165 | 4.9367 | 0.032075 | 271 | knn | NumNeighbors: 2131 | | 17 | 6 | Accept | 0.038998 | 1.3761 | 0.032075 | 271 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 3.2054e-06 | | | | | | | | | | LayerSizes: 74 | | 18 | 6 | Accept | 0.12327 | 1.8297 | 0.032075 | 271 | knn | NumNeighbors: 34 | | 19 | 6 | Accept | 0.73929 | 12.116 | 0.032075 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 91.704 | | | | | | | | | | KernelScale: 0.015616 | | 20 | 6 | Accept | 0.17634 | 3.8994 | 0.032075 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.003312 | | | | | | | | | | KernelScale: 851.79 | |====================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Training set | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | size | | | |====================================================================================================================================================| | 21 | 6 | Accept | 0.74165 | 4.6497 | 0.032075 | 271 | knn | NumNeighbors: 678 | | 22 | 6 | Accept | 0.13587 | 2.3404 | 0.032075 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.019161 | | | | | | | | | | KernelScale: 153.3 | | 23 | 6 | Best | 0.026352 | 5.17 | 0.026352 | 1084 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 3.2054e-06 | | | | | | | | | | LayerSizes: 74 | | 24 | 6 | Accept | 0.74165 | 10.604 | 0.026352 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 0.0070102 | | | | | | | | | | KernelScale: 0.0038694 | | 25 | 6 | Accept | 0.043243 | 2.9641 | 0.026352 | 1084 | knn | NumNeighbors: 2 | | 26 | 6 | Best | 0.011861 | 29.125 | 0.011861 | 4334 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 3.2054e-06 | | | | | | | | | | LayerSizes: 74 | | 27 | 6 | Accept | 0.63379 | 66.627 | 0.011861 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 0.0028215 | | | | | | | | | | Standardize: true | | 28 | 6 | Accept | 0.74165 | 4.5204 | 0.011861 | 271 | knn | NumNeighbors: 1428 | | 29 | 6 | Accept | 0.05755 | 1.7987 | 0.011861 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 1.0426 | | | | | | | | | | KernelScale: 15.383 | | 30 | 6 | Accept | 0.054458 | 0.67873 | 0.011861 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 31 | 6 | Accept | 0.053904 | 0.66551 | 0.011861 | 1084 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 32 | 6 | Accept | 0.55727 | 91.003 | 0.011861 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 0.0059208 | | | | | | | | | | Standardize: false | | 33 | 6 | Accept | 0.74165 | 11.662 | 0.011861 | 271 | ensemble | Method: AdaBoostM2 | | | | | | | | | | NumLearningCycles: 270 | | | | | | | | | | MinLeafSize: 1014 | | | | | | | | | | MaxNumSplits: 27 | | 34 | 6 | Accept | 0.74165 | 10.501 | 0.011861 | 271 | ensemble | Method: AdaBoostM2 | | | | | | | | | | NumLearningCycles: 300 | | | | | | | | | | MinLeafSize: 184 | | | | | | | | | | MaxNumSplits: 18 | | 35 | 6 | Accept | 0.73842 | 168.5 | 0.011861 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 17.493 | | | | | | | | | | Standardize: true | | 36 | 6 | Accept | 0.73417 | 171.15 | 0.011861 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 856.62 | | | | | | | | | | Standardize: false | | 37 | 6 | Accept | 0.19139 | 173.58 | 0.011861 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 4.6757 | | | | | | | | | | Standardize: true | | 38 | 6 | Accept | 0.032536 | 3.7049 | 0.011861 | 1084 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 1.0426 | | | | | | | | | | KernelScale: 15.383 | | 39 | 6 | Accept | 0.74165 | 1.524 | 0.011861 | 271 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.29242 | | | | | | | | | | LayerSizes: [ 20 14 ] | | 40 | 6 | Accept | 0.052381 | 1.2116 | 0.011861 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | |====================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Training set | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | size | | | |====================================================================================================================================================| | 41 | 6 | Accept | 0.054966 | 0.65989 | 0.011861 | 1084 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 42 | 6 | Accept | 0.029952 | 18.516 | 0.011861 | 271 | ensemble | Method: AdaBoostM2 | | | | | | | | | | NumLearningCycles: 229 | | | | | | | | | | MinLeafSize: 7 | | | | | | | | | | MaxNumSplits: 31 | | 43 | 6 | Accept | 0.62415 | 4.6925 | 0.011861 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.019139 | | | | | | | | | | KernelScale: 0.86008 | | 44 | 6 | Accept | 0.069503 | 32.867 | 0.011861 | 271 | net | Activations: tanh | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.096e-06 | | | | | | | | | | LayerSizes: [ 289 2 ] | | 45 | 6 | Accept | 0.051366 | 1.214 | 0.011861 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 46 | 6 | Accept | 0.73888 | 11.397 | 0.011861 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 939.27 | | | | | | | | | | KernelScale: 0.0013942 | | 47 | 6 | Accept | 0.027691 | 34.472 | 0.011861 | 4334 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.0016149 | | | | | | | | | | LayerSizes: 18 | | 48 | 6 | Accept | 0.15341 | 0.51534 | 0.011861 | 271 | tree | MinLeafSize: 58 | | 49 | 6 | Accept | 0.32204 | 4.652 | 0.011861 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 4.6764 | | | | | | | | | | KernelScale: 2.4604 | | 50 | 6 | Accept | 0.11459 | 2.3006 | 0.011861 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 94.31 | | | | | | | | | | KernelScale: 944.78 | | 51 | 6 | Accept | 0.047351 | 0.96901 | 0.011861 | 1084 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 52 | 6 | Accept | 0.04412 | 0.66376 | 0.011861 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 53 | 6 | Accept | 0.048689 | 1.4012 | 0.011861 | 271 | knn | NumNeighbors: 1 | | 54 | 6 | Accept | 0.05492 | 0.73525 | 0.011861 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 55 | 6 | Accept | 0.73569 | 9.3198 | 0.011861 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.0031837 | | | | | | | | | | KernelScale: 0.29449 | | 56 | 6 | Accept | 0.053397 | 0.78515 | 0.011861 | 1084 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 57 | 6 | Accept | 0.11847 | 167.45 | 0.011861 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 2.7531 | | | | | | | | | | Standardize: true | | 58 | 6 | Accept | 0.049382 | 0.82672 | 0.011861 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 59 | 6 | Accept | 0.064242 | 1.1364 | 0.011861 | 271 | knn | NumNeighbors: 2 | | 60 | 6 | Accept | 0.076565 | 0.88796 | 0.011861 | 271 | tree | MinLeafSize: 2 | |====================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Training set | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | size | | | |====================================================================================================================================================| | 61 | 6 | Accept | 0.059904 | 0.94055 | 0.011861 | 271 | knn | NumNeighbors: 1 | | 62 | 6 | Accept | 0.033875 | 3.2696 | 0.011861 | 1084 | knn | NumNeighbors: 1 | | 63 | 6 | Accept | 0.066181 | 9.5233 | 0.011861 | 271 | net | Activations: relu | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 3.0708e-06 | | | | | | | | | | LayerSizes: [ 85 6 13 ] | | 64 | 6 | Accept | 0.71045 | 6.4658 | 0.011861 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 86.648 | | | | | | | | | | KernelScale: 0.68451 | | 65 | 6 | Accept | 0.015368 | 27.396 | 0.011861 | 1084 | ensemble | Method: AdaBoostM2 | | | | | | | | | | NumLearningCycles: 229 | | | | | | | | | | MinLeafSize: 7 | | | | | | | | | | MaxNumSplits: 31 | | 66 | 6 | Accept | 0.73906 | 11.212 | 0.011861 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 1.622 | | | | | | | | | | KernelScale: 0.0038764 | | 67 | 6 | Accept | 0.053904 | 0.66704 | 0.011861 | 1084 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 68 | 6 | Accept | 0.73911 | 11.194 | 0.011861 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 49.928 | | | | | | | | | | KernelScale: 0.0035535 | | 69 | 6 | Accept | 0.74165 | 0.19475 | 0.011861 | 271 | tree | MinLeafSize: 1365 | | 70 | 6 | Accept | 0.046751 | 3.2493 | 0.011861 | 271 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 1.0454e-06 | | | | | | | | | | LayerSizes: [ 4 7 26 ] | | 71 | 6 | Best | 0.0077995 | 37.224 | 0.0077995 | 4334 | ensemble | Method: AdaBoostM2 | | | | | | | | | | NumLearningCycles: 229 | | | | | | | | | | MinLeafSize: 7 | | | | | | | | | | MaxNumSplits: 31 | | 72 | 6 | Accept | 0.096963 | 3.5085 | 0.0077995 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 8.0828 | | | | | | | | | | KernelScale: 171.49 | | 73 | 6 | Accept | 0.74165 | 168.14 | 0.0077995 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 596 | | | | | | | | | | Standardize: true | | 74 | 6 | Accept | 0.057227 | 0.76102 | 0.0077995 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 75 | 6 | Accept | 0.059304 | 14.374 | 0.0077995 | 1084 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 1.0454e-06 | | | | | | | | | | LayerSizes: [ 4 7 26 ] | | 76 | 6 | Accept | 0.73975 | 11.338 | 0.0077995 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.56028 | | | | | | | | | | KernelScale: 0.0093903 | | 77 | 6 | Accept | 0.057966 | 0.71414 | 0.0077995 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 78 | 6 | Accept | 0.046151 | 0.7308 | 0.0077995 | 1084 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 79 | 6 | Accept | 0.048505 | 173.22 | 0.0077995 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 0.61252 | | | | | | | | | | Standardize: true | | 80 | 6 | Accept | 0.068257 | 20.884 | 0.0077995 | 271 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.0891e-05 | | | | | | | | | | LayerSizes: [ 80 86 2 ] | |====================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Training set | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | size | | | |====================================================================================================================================================| | 81 | 6 | Accept | 0.74165 | 4.8285 | 0.0077995 | 271 | knn | NumNeighbors: 1560 | | 82 | 6 | Accept | 0.72946 | 168.86 | 0.0077995 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 754.46 | | | | | | | | | | Standardize: false | | 83 | 6 | Accept | 0.3807 | 169.46 | 0.0077995 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 44.255 | | | | | | | | | | Standardize: false | | 84 | 6 | Accept | 0.081549 | 130.35 | 0.0077995 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 0.098769 | | | | | | | | | | Standardize: true | | 85 | 6 | Accept | 0.74165 | 3.7268 | 0.0077995 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 8.8662 | | | | | | | | | | KernelScale: 541.83 | | 86 | 6 | Accept | 0.10033 | 0.66666 | 0.0077995 | 271 | tree | MinLeafSize: 17 | | 87 | 6 | Accept | 0.25448 | 171.99 | 0.0077995 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 7.1617 | | | | | | | | | | Standardize: true | | 88 | 6 | Accept | 0.05755 | 0.71855 | 0.0077995 | 1084 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 89 | 6 | Accept | 0.049105 | 1.283 | 0.0077995 | 271 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 4.2009e-08 | | | | | | | | | | LayerSizes: 133 | | 90 | 6 | Accept | 0.072457 | 0.73998 | 0.0077995 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 91 | 6 | Accept | 0.021922 | 6.709 | 0.0077995 | 4334 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 1.0426 | | | | | | | | | | KernelScale: 15.383 | | 92 | 6 | Accept | 0.73883 | 12.629 | 0.0077995 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.19021 | | | | | | | | | | KernelScale: 0.13128 | | 93 | 6 | Accept | 0.74165 | 10.879 | 0.0077995 | 271 | ensemble | Method: AdaBoostM2 | | | | | | | | | | NumLearningCycles: 288 | | | | | | | | | | MinLeafSize: 4760 | | | | | | | | | | MaxNumSplits: 25 | | 94 | 6 | Accept | 0.030967 | 3.5848 | 0.0077995 | 1084 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 4.2009e-08 | | | | | | | | | | LayerSizes: 133 | | 95 | 6 | Accept | 0.067842 | 0.71485 | 0.0077995 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 96 | 6 | Accept | 0.74165 | 0.21973 | 0.0077995 | 271 | tree | MinLeafSize: 5731 | | 97 | 6 | Accept | 0.51435 | 97.511 | 0.0077995 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 0.010292 | | | | | | | | | | Standardize: false | | 98 | 6 | Accept | 0.049474 | 0.7033 | 0.0077995 | 1084 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 99 | 6 | Accept | 0.078503 | 3.5346 | 0.0077995 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 65.734 | | | | | | | | | | KernelScale: 4.5947 | | 100 | 6 | Accept | 0.047628 | 0.73303 | 0.0077995 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | |====================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Training set | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | size | | | |====================================================================================================================================================| | 101 | 6 | Accept | 0.037428 | 7.6392 | 0.0077995 | 271 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 6.2668e-05 | | | | | | | | | | LayerSizes: [ 171 9 ] | | 102 | 6 | Accept | 0.73962 | 9.4795 | 0.0077995 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 28.585 | | | | | | | | | | KernelScale: 0.14053 | | 103 | 6 | Accept | 0.74165 | 164.68 | 0.0077995 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 209.59 | | | | | | | | | | Standardize: true | | 104 | 6 | Accept | 0.026814 | 28.779 | 0.0077995 | 1084 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 6.2668e-05 | | | | | | | | | | LayerSizes: [ 171 9 ] | | 105 | 6 | Best | 0.0053074 | 62.586 | 0.0053074 | 17335 | ensemble | Method: AdaBoostM2 | | | | | | | | | | NumLearningCycles: 229 | | | | | | | | | | MinLeafSize: 7 | | | | | | | | | | MaxNumSplits: 31 | | 106 | 6 | Accept | 0.74792 | 1.5953 | 0.0053074 | 271 | ensemble | Method: AdaBoostM2 | | | | | | | | | | NumLearningCycles: 238 | | | | | | | | | | MinLeafSize: 1 | | | | | | | | | | MaxNumSplits: 57 | | 107 | 6 | Accept | 0.74165 | 0.3562 | 0.0053074 | 271 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.6829 | | | | | | | | | | LayerSizes: [ 40 4 ] | | 108 | 6 | Accept | 0.060227 | 3.6079 | 0.0053074 | 271 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.0069133 | | | | | | | | | | LayerSizes: [ 3 12 12 ] | | 109 | 6 | Accept | 0.049658 | 0.70789 | 0.0053074 | 1084 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 110 | 6 | Accept | 0.59156 | 79.711 | 0.0053074 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 0.0033878 | | | | | | | | | | Standardize: false | | 111 | 6 | Accept | 0.74165 | 5.2086 | 0.0053074 | 271 | knn | NumNeighbors: 373 | | 112 | 6 | Accept | 0.047766 | 0.83263 | 0.0053074 | 271 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 2.319e-05 | | | | | | | | | | LayerSizes: 17 | | 113 | 6 | Accept | 0.054089 | 0.72423 | 0.0053074 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 114 | 6 | Accept | 0.02866 | 3.1959 | 0.0053074 | 1084 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 2.319e-05 | | | | | | | | | | LayerSizes: 17 | | 115 | 6 | Accept | 0.74165 | 4.4465 | 0.0053074 | 271 | knn | NumNeighbors: 1189 | | 116 | 6 | Accept | 0.74165 | 4.442 | 0.0053074 | 271 | knn | NumNeighbors: 1145 | | 117 | 6 | Accept | 0.032444 | 18.279 | 0.0053074 | 271 | ensemble | Method: Bag | | | | | | | | | | NumLearningCycles: 268 | | | | | | | | | | MinLeafSize: 1 | | | | | | | | | | MaxNumSplits: 79 | | 118 | 6 | Accept | 0.069919 | 0.70972 | 0.0053074 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 119 | 6 | Accept | 0.098025 | 155.9 | 0.0053074 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 6.0379 | | | | | | | | | | Standardize: false | | 120 | 6 | Accept | 0.14745 | 161.41 | 0.0053074 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 2.9595 | | | | | | | | | | Standardize: true | |====================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Training set | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | size | | | |====================================================================================================================================================| | 121 | 6 | Accept | 0.046243 | 0.6139 | 0.0053074 | 271 | net | Activations: relu | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 6.6757e-07 | | | | | | | | | | LayerSizes: 65 | | 122 | 6 | Accept | 0.053766 | 2.3698 | 0.0053074 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 914.18 | | | | | | | | | | KernelScale: 85.472 | | 123 | 6 | Accept | 0.025752 | 1.2539 | 0.0053074 | 1084 | net | Activations: relu | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 6.6757e-07 | | | | | | | | | | LayerSizes: 65 | | 124 | 6 | Accept | 0.73925 | 10.728 | 0.0053074 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 544.31 | | | | | | | | | | KernelScale: 0.007687 | | 125 | 6 | Accept | 0.054966 | 14.621 | 0.0053074 | 271 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.9041e-06 | | | | | | | | | | LayerSizes: [ 210 9 ] | | 126 | 6 | Accept | 0.02806 | 25.852 | 0.0053074 | 1084 | ensemble | Method: Bag | | | | | | | | | | NumLearningCycles: 268 | | | | | | | | | | MinLeafSize: 1 | | | | | | | | | | MaxNumSplits: 79 | | 127 | 6 | Accept | 0.04412 | 1.0243 | 0.0053074 | 271 | tree | MinLeafSize: 2 | | 128 | 6 | Accept | 0.74165 | 4.9624 | 0.0053074 | 271 | knn | NumNeighbors: 2336 | | 129 | 6 | Accept | 0.12244 | 2.6488 | 0.0053074 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.045281 | | | | | | | | | | KernelScale: 593.59 | | 130 | 6 | Accept | 0.030321 | 15.349 | 0.0053074 | 271 | ensemble | Method: Bag | | | | | | | | | | NumLearningCycles: 205 | | | | | | | | | | MinLeafSize: 1 | | | | | | | | | | MaxNumSplits: 76 | | 131 | 5 | Accept | 0.077441 | 1.2571 | 0.0053074 | 271 | knn | NumNeighbors: 6 | | 132 | 5 | Accept | 0.065811 | 0.75844 | 0.0053074 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 133 | 6 | Accept | 0.05492 | 2.1752 | 0.0053074 | 1084 | tree | MinLeafSize: 2 | | 134 | 6 | Accept | 0.068303 | 1.1461 | 0.0053074 | 271 | knn | NumNeighbors: 6 | | 135 | 6 | Accept | 0.74165 | 9.9296 | 0.0053074 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 0.012836 | | | | | | | | | | KernelScale: 0.0013573 | | 136 | 6 | Accept | 0.048182 | 3.954 | 0.0053074 | 271 | net | Activations: tanh | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.0917e-09 | | | | | | | | | | LayerSizes: 27 | | 137 | 6 | Accept | 0.74165 | 4.8146 | 0.0053074 | 271 | knn | NumNeighbors: 5959 | | 138 | 6 | Accept | 0.027368 | 20.876 | 0.0053074 | 1084 | ensemble | Method: Bag | | | | | | | | | | NumLearningCycles: 205 | | | | | | | | | | MinLeafSize: 1 | | | | | | | | | | MaxNumSplits: 76 | | 139 | 6 | Accept | 0.037936 | 1.6741 | 0.0053074 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 3.9775 | | | | | | | | | | KernelScale: 13.747 | | 140 | 6 | Accept | 0.10181 | 2.2241 | 0.0053074 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.5329 | | | | | | | | | | KernelScale: 458.88 | |====================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Training set | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | size | | | |====================================================================================================================================================| | 141 | 6 | Accept | 0.066688 | 164.37 | 0.0053074 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 1.1777 | | | | | | | | | | Standardize: true | | 142 | 6 | Accept | 0.049474 | 14.68 | 0.0053074 | 1084 | net | Activations: tanh | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.0917e-09 | | | | | | | | | | LayerSizes: 27 | | 143 | 6 | Accept | 0.076611 | 8.3053 | 0.0053074 | 271 | net | Activations: relu | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.2759e-05 | | | | | | | | | | LayerSizes: [ 71 14 ] | | 144 | 6 | Accept | 0.012322 | 39.233 | 0.0053074 | 4334 | net | Activations: relu | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 6.6757e-07 | | | | | | | | | | LayerSizes: 65 | | 145 | 6 | Accept | 0.74165 | 4.99 | 0.0053074 | 271 | knn | NumNeighbors: 980 | | 146 | 6 | Accept | 0.026445 | 2.363 | 0.0053074 | 1084 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 3.9775 | | | | | | | | | | KernelScale: 13.747 | | 147 | 6 | Accept | 0.04209 | 2.3578 | 0.0053074 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 10.427 | | | | | | | | | | KernelScale: 16.103 | | 148 | 6 | Accept | 0.069688 | 0.70265 | 0.0053074 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 149 | 6 | Accept | 0.74165 | 4.537 | 0.0053074 | 271 | knn | NumNeighbors: 767 | | 150 | 6 | Accept | 0.023214 | 2.3517 | 0.0053074 | 1084 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 10.427 | | | | | | | | | | KernelScale: 16.103 | | 151 | 6 | Accept | 0.73929 | 9.8139 | 0.0053074 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 6.3626 | | | | | | | | | | KernelScale: 0.002014 | | 152 | 6 | Accept | 0.011215 | 176.8 | 0.0053074 | 4334 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 6.2668e-05 | | | | | | | | | | LayerSizes: [ 171 9 ] | | 153 | 6 | Accept | 0.011769 | 3.7526 | 0.0053074 | 4334 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 10.427 | | | | | | | | | | KernelScale: 16.103 | | 154 | 6 | Accept | 0.74165 | 0.17534 | 0.0053074 | 271 | tree | MinLeafSize: 3138 | | 155 | 6 | Accept | 0.74165 | 10.965 | 0.0053074 | 271 | ensemble | Method: Bag | | | | | | | | | | NumLearningCycles: 265 | | | | | | | | | | MinLeafSize: 272 | | | | | | | | | | MaxNumSplits: 32 | | 156 | 6 | Accept | 0.74165 | 9.6757 | 0.0053074 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 0.0027945 | | | | | | | | | | KernelScale: 0.047512 | | 157 | 6 | Accept | 0.055058 | 11.792 | 0.0053074 | 271 | net | Activations: relu | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.0004489 | | | | | | | | | | LayerSizes: [ 127 10 ] | | 158 | 6 | Accept | 0.028014 | 2.2151 | 0.0053074 | 1084 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 914.18 | | | | | | | | | | KernelScale: 85.472 | | 159 | 6 | Accept | 0.044167 | 0.74389 | 0.0053074 | 271 | net | Activations: relu | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 1.267e-06 | | | | | | | | | | LayerSizes: [ 21 51 ] | | 160 | 6 | Accept | 0.74165 | 4.4576 | 0.0053074 | 271 | knn | NumNeighbors: 1090 | |====================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Training set | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | size | | | |====================================================================================================================================================| | 161 | 6 | Accept | 0.40027 | 4.8502 | 0.0053074 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 24.409 | | | | | | | | | | KernelScale: 1.6395 | | 162 | 6 | Accept | 0.022199 | 1.9746 | 0.0053074 | 1084 | net | Activations: relu | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 1.267e-06 | | | | | | | | | | LayerSizes: [ 21 51 ] | | 163 | 6 | Accept | 0.043013 | 292.66 | 0.0053074 | 1084 | nb | DistributionNames: kernel | | | | | | | | | | Width: 0.61252 | | | | | | | | | | Standardize: true | | 164 | 6 | Accept | 0.74165 | 8.3033 | 0.0053074 | 271 | ensemble | Method: Bag | | | | | | | | | | NumLearningCycles: 216 | | | | | | | | | | MinLeafSize: 8895 | | | | | | | | | | MaxNumSplits: 54 | | 165 | 6 | Accept | 0.060412 | 0.74162 | 0.0053074 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 166 | 6 | Accept | 0.74165 | 4.6892 | 0.0053074 | 271 | knn | NumNeighbors: 4825 | | 167 | 6 | Accept | 0.053812 | 1.6486 | 0.0053074 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 41.59 | | | | | | | | | | KernelScale: 66.838 | | 168 | 6 | Accept | 0.73934 | 9.6965 | 0.0053074 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 3.5182 | | | | | | | | | | KernelScale: 0.0021656 | | 169 | 6 | Accept | 0.040844 | 0.65233 | 0.0053074 | 271 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 3.4639e-08 | | | | | | | | | | LayerSizes: 12 | | 170 | 6 | Accept | 0.65807 | 0.50924 | 0.0053074 | 271 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.2677 | | | | | | | | | | LayerSizes: 46 | | 171 | 6 | Accept | 0.028844 | 2.7895 | 0.0053074 | 1084 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 41.59 | | | | | | | | | | KernelScale: 66.838 | | 172 | 6 | Accept | 0.041213 | 2.536 | 0.0053074 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 114.59 | | | | | | | | | | KernelScale: 169.23 | | 173 | 6 | Accept | 0.064981 | 7.9256 | 0.0053074 | 271 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 6.46e-10 | | | | | | | | | | LayerSizes: [ 94 6 ] | | 174 | 6 | Accept | 0.033459 | 1.8565 | 0.0053074 | 1084 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 3.4639e-08 | | | | | | | | | | LayerSizes: 12 | | 175 | 6 | Accept | 0.059904 | 0.80685 | 0.0053074 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 176 | 6 | Accept | 0.28761 | 1.5076 | 0.0053074 | 271 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 9.7994e-09 | | | | | | | | | | LayerSizes: 1 | | 177 | 6 | Accept | 0.73957 | 11.525 | 0.0053074 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.01816 | | | | | | | | | | KernelScale: 0.0019383 | | 178 | 6 | Accept | 0.1319 | 2.3554 | 0.0053074 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.065299 | | | | | | | | | | KernelScale: 930.44 | | 179 | 6 | Accept | 0.027829 | 2.7788 | 0.0053074 | 1084 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 114.59 | | | | | | | | | | KernelScale: 169.23 | | 180 | 6 | Accept | 0.07338 | 0.72933 | 0.0053074 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | |====================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Training set | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | size | | | |====================================================================================================================================================| | 181 | 6 | Accept | 0.74165 | 4.4535 | 0.0053074 | 271 | knn | NumNeighbors: 232 | | 182 | 6 | Accept | 0.059119 | 0.77186 | 0.0053074 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 183 | 6 | Accept | 0.050535 | 0.69862 | 0.0053074 | 1084 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 184 | 6 | Accept | 0.74165 | 10.098 | 0.0053074 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 0.0028005 | | | | | | | | | | KernelScale: 0.010353 | | 185 | 6 | Accept | 0.046243 | 0.71168 | 0.0053074 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 186 | 6 | Accept | 0.74165 | 4.6281 | 0.0053074 | 271 | knn | NumNeighbors: 857 | | 187 | 6 | Accept | 0.1817 | 2.3545 | 0.0053074 | 271 | knn | NumNeighbors: 66 | | 188 | 6 | Accept | 0.051551 | 0.70548 | 0.0053074 | 1084 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 189 | 6 | Accept | 0.74165 | 4.6952 | 0.0053074 | 271 | knn | NumNeighbors: 6918 | | 190 | 6 | Accept | 0.04329 | 1.515 | 0.0053074 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 900.92 | | | | | | | | | | KernelScale: 44.444 | | 191 | 6 | Accept | 0.72199 | 6.2291 | 0.0053074 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 28.613 | | | | | | | | | | KernelScale: 0.4911 | | 192 | 6 | Accept | 0.043936 | 78.329 | 0.0053074 | 271 | ensemble | Coding: onevsone | | | | | | | | | | Method: LogitBoost | | | | | | | | | | NumLearningCycles: 211 | | | | | | | | | | MinLeafSize: 4 | | | | | | | | | | MaxNumSplits: 13 | | 193 | 6 | Accept | 0.036275 | 0.85738 | 0.0053074 | 271 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 5.0779e-06 | | | | | | | | | | LayerSizes: 31 | | 194 | 6 | Accept | 0.014353 | 6.0141 | 0.0053074 | 4334 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 3.9775 | | | | | | | | | | KernelScale: 13.747 | | 195 | 6 | Accept | 0.039921 | 0.67116 | 0.0053074 | 271 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.00035338 | | | | | | | | | | LayerSizes: 4 | | 196 | 6 | Accept | 0.027552 | 2.0495 | 0.0053074 | 1084 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 900.92 | | | | | | | | | | KernelScale: 44.444 | | 197 | 6 | Accept | 0.52917 | 0.28918 | 0.0053074 | 271 | tree | MinLeafSize: 116 | | 198 | 6 | Accept | 0.083718 | 1.4053 | 0.0053074 | 271 | knn | NumNeighbors: 12 | | 199 | 6 | Accept | 0.74165 | 0.63448 | 0.0053074 | 271 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.27358 | | | | | | | | | | LayerSizes: [ 21 58 47 ] | | 200 | 6 | Accept | 0.068903 | 2.0491 | 0.0053074 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.53387 | | | | | | | | | | KernelScale: 9.7797 | |====================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Training set | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | size | | | |====================================================================================================================================================| | 201 | 6 | Accept | 0.023675 | 2.9905 | 0.0053074 | 1084 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 5.0779e-06 | | | | | | | | | | LayerSizes: 31 | | 202 | 6 | Accept | 0.74165 | 81.003 | 0.0053074 | 271 | ensemble | Coding: onevsone | | | | | | | | | | Method: LogitBoost | | | | | | | | | | NumLearningCycles: 242 | | | | | | | | | | MinLeafSize: 2550 | | | | | | | | | | MaxNumSplits: 42 | | 203 | 6 | Accept | 0.74165 | 4.7264 | 0.0053074 | 271 | knn | NumNeighbors: 911 | | 204 | 6 | Accept | 0.031706 | 2.5716 | 0.0053074 | 1084 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.00035338 | | | | | | | | | | LayerSizes: 4 | | 205 | 6 | Accept | 0.058012 | 0.75697 | 0.0053074 | 271 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 206 | 6 | Accept | 0.013338 | 54.224 | 0.0053074 | 4334 | net | Activations: relu | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 1.267e-06 | | | | | | | | | | LayerSizes: [ 21 51 ] | | 207 | 6 | Accept | 0.74165 | 0.54395 | 0.0053074 | 271 | net | Activations: tanh | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.40394 | | | | | | | | | | LayerSizes: [ 23 93 ] | | 208 | 6 | Accept | 0.10167 | 2.3728 | 0.0053074 | 271 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 3.9533e-09 | | | | | | | | | | LayerSizes: [ 1 1 ] | | 209 | 6 | Accept | 0.74165 | 4.6078 | 0.0053074 | 271 | knn | NumNeighbors: 862 | | 210 | 6 | Accept | 0.36815 | 3.3522 | 0.0053074 | 271 | knn | NumNeighbors: 140 | | 211 | 6 | Accept | 0.74165 | 4.4738 | 0.0053074 | 271 | knn | NumNeighbors: 8006 | | 212 | 6 | Accept | 0.044997 | 43.489 | 0.0053074 | 271 | net | Activations: tanh | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 2.5417e-08 | | | | | | | | | | LayerSizes: [ 44 116 226 ] | | 213 | 6 | Accept | 0.05132 | 3.4248 | 0.0053074 | 271 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.0027776 | | | | | | | | | | LayerSizes: [ 72 8 ] | | 214 | 6 | Accept | 0.74165 | 45.286 | 0.0053074 | 271 | ensemble | Coding: onevsall | | | | | | | | | | Method: LogitBoost | | | | | | | | | | NumLearningCycles: 268 | | | | | | | | | | MinLeafSize: 2870 | | | | | | | | | | MaxNumSplits: 91 | | 215 | 6 | Accept | 0.7398 | 9.7726 | 0.0053074 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 68.727 | | | | | | | | | | KernelScale: 0.025309 | | 216 | 6 | Accept | 0.74165 | 4.64 | 0.0053074 | 271 | knn | NumNeighbors: 4617 | | 217 | 6 | Accept | 0.66781 | 4.5932 | 0.0053074 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.45784 | | | | | | | | | | KernelScale: 0.67635 | | 218 | 6 | Accept | 0.046843 | 5.8551 | 0.0053074 | 1084 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.0027776 | | | | | | | | | | LayerSizes: [ 72 8 ] | | 219 | 6 | Accept | 0.013153 | 26.521 | 0.0053074 | 4334 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 5.0779e-06 | | | | | | | | | | LayerSizes: 31 | | 220 | 6 | Accept | 0.74165 | 0.13889 | 0.0053074 | 271 | tree | MinLeafSize: 214 | |====================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Training set | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | size | | | |====================================================================================================================================================| | 221 | 6 | Accept | 0.042828 | 3.9304 | 0.0053074 | 271 | net | Activations: tanh | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.00032581 | | | | | | | | | | LayerSizes: 10 | | 222 | 6 | Accept | 0.74165 | 9.8725 | 0.0053074 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 0.32491 | | | | | | | | | | KernelScale: 0.0011646 | | 223 | 6 | Accept | 0.74165 | 0.12981 | 0.0053074 | 271 | tree | MinLeafSize: 405 | | 224 | 6 | Accept | 0.036829 | 9.849 | 0.0053074 | 1084 | net | Activations: tanh | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.00032581 | | | | | | | | | | LayerSizes: 10 | | 225 | 6 | Accept | 0.10001 | 1.5959 | 0.0053074 | 271 | knn | NumNeighbors: 24 | | 226 | 6 | Accept | 0.11805 | 0.42103 | 0.0053074 | 271 | tree | MinLeafSize: 51 | | 227 | 6 | Accept | 0.029444 | 93.524 | 0.0053074 | 1084 | ensemble | Coding: onevsone | | | | | | | | | | Method: LogitBoost | | | | | | | | | | NumLearningCycles: 211 | | | | | | | | | | MinLeafSize: 4 | | | | | | | | | | MaxNumSplits: 13 | | 228 | 6 | Accept | 0.74165 | 0.19998 | 0.0053074 | 271 | tree | MinLeafSize: 2088 | | 229 | 6 | Accept | 0.7416 | 168.62 | 0.0053074 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 20.609 | | | | | | | | | | Standardize: true | | 230 | 6 | Accept | 0.032906 | 151.18 | 0.0053074 | 1084 | net | Activations: tanh | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 2.5417e-08 | | | | | | | | | | LayerSizes: [ 44 116 226 ] | | 231 | 6 | Accept | 0.6787 | 70.071 | 0.0053074 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 0.0015605 | | | | | | | | | | Standardize: false | | 232 | 6 | Accept | 0.12955 | 3.7371 | 0.0053074 | 271 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 2.7261e-07 | | | | | | | | | | LayerSizes: 5 | | 233 | 6 | Accept | 0.52838 | 0.89457 | 0.0053074 | 271 | tree | MinLeafSize: 105 | | 234 | 6 | Accept | 0.055474 | 0.95035 | 0.0053074 | 271 | knn | NumNeighbors: 3 | | 235 | 6 | Accept | 0.74165 | 9.9179 | 0.0053074 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 0.2143 | | | | | | | | | | KernelScale: 0.19817 | | 236 | 6 | Accept | 0.032029 | 35.514 | 0.0053074 | 1084 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.9041e-06 | | | | | | | | | | LayerSizes: [ 210 9 ] | | 237 | 6 | Accept | 0.34696 | 8.8977 | 0.0053074 | 271 | net | Activations: relu | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.0022429 | | | | | | | | | | LayerSizes: [ 100 1 ] | | 238 | 6 | Accept | 0.28452 | 95.413 | 0.0053074 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 0.016256 | | | | | | | | | | Standardize: true | | 239 | 6 | Accept | 0.74165 | 171.77 | 0.0053074 | 271 | nb | DistributionNames: kernel | | | | | | | | | | Width: 132.12 | | | | | | | | | | Standardize: true | | 240 | 6 | Accept | 0.72877 | 7.0112 | 0.0053074 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 80.2 | | | | | | | | | | KernelScale: 0.4411 | |====================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Training set | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | size | | | |====================================================================================================================================================| | 241 | 6 | Accept | 0.03429 | 3.2421 | 0.0053074 | 1084 | knn | NumNeighbors: 3 | | 242 | 6 | Accept | 0.13144 | 8.5231 | 0.0053074 | 271 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 4.1361e-06 | | | | | | | | | | LayerSizes: [ 1 102 ] | | 243 | 6 | Accept | 0.058058 | 4.1342 | 0.0053074 | 271 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.00060115 | | | | | | | | | | LayerSizes: [ 3 8 ] | | 244 | 6 | Accept | 0.036598 | 33.658 | 0.0053074 | 1084 | net | Activations: relu | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.0004489 | | | | | | | | | | LayerSizes: [ 127 10 ] | | 245 | 6 | Accept | 0.12756 | 3.8626 | 0.0053074 | 271 | net | Activations: tanh | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.0034719 | | | | | | | | | | LayerSizes: [ 2 10 ] | | 246 | 6 | Accept | 0.05012 | 0.98248 | 0.0053074 | 1084 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 247 | 6 | Accept | 0.064842 | 0.86246 | 0.0053074 | 271 | tree | MinLeafSize: 6 | | 248 | 6 | Accept | 0.32126 | 16.422 | 0.0053074 | 271 | ensemble | Method: Bag | | | | | | | | | | NumLearningCycles: 287 | | | | | | | | | | MinLeafSize: 92 | | | | | | | | | | MaxNumSplits: 34 | | 249 | 6 | Accept | 0.74165 | 3.7533 | 0.0053074 | 271 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 0.0020576 | | | | | | | | | | KernelScale: 601.64 | | 250 | 6 | Accept | 0.73985 | 10.991 | 0.0053074 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.021505 | | | | | | | | | | KernelScale: 0.0044067 | | 251 | 6 | Accept | 0.01846 | 46.153 | 0.0053074 | 4334 | ensemble | Method: Bag | | | | | | | | | | NumLearningCycles: 205 | | | | | | | | | | MinLeafSize: 1 | | | | | | | | | | MaxNumSplits: 76 | | 252 | 6 | Accept | 0.12936 | 8.0856 | 0.0053074 | 1084 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.00060115 | | | | | | | | | | LayerSizes: [ 3 8 ] | | 253 | 6 | Accept | 0.013707 | 10.948 | 0.0053074 | 4334 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 900.92 | | | | | | | | | | KernelScale: 44.444 | | 254 | 6 | Accept | 0.30598 | 20.762 | 0.0053074 | 271 | net | Activations: tanh | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.5659e-09 | | | | | | | | | | LayerSizes: [ 173 2 ] | | 255 | 6 | Accept | 0.48768 | 4.4372 | 0.0053074 | 271 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 3.1297 | | | | | | | | | | KernelScale: 1.1348 | | 256 | 6 | Accept | 0.006415 | 11.83 | 0.0053074 | 17335 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 10.427 | | | | | | | | | | KernelScale: 16.103 | | 257 | 6 | Accept | 0.064427 | 0.76725 | 0.0053074 | 271 | nb | DistributionNames: ...

__________________________________________________________ Optimization completed. Total iterations: 595 Total elapsed time: 2507.5379 seconds Total time for training and validation: 14243.9529 seconds Best observed learner is a net model with: Learner: net Activations: sigmoid Standardize: true Lambda: 3.2054e-06 LayerSizes: 74 Observed validation loss: 0.0043843 Time for training and validation: 193.32 seconds Documentation for fitcauto display
The final model returned by fitcauto corresponds to the best observed learner. Before returning the model, the function retrains it using all the training data (XTrain and YTrain), the listed Learner (or model) type, and the displayed hyperparameter values.
Evaluate Test Set Performance
Evaluate the final model performance on the test data set.
testAccuracy = 1 - loss(Mdl,XTest,YTest)
testAccuracy = 0.9992
The final model correctly classifies over 99% of the observations.
Use fitcauto to automatically select a classification model with optimized hyperparameters, given predictor and response data stored in a table. Before passing data to fitcauto, perform feature selection to remove unimportant predictors from the data set.
Load and Partition Data
Read the sample file CreditRating_Historical.dat into a table. The predictor data consists of financial ratios and industry sector information for a list of corporate customers. The response variable consists of credit ratings assigned by a rating agency. Preview the first few rows of the data set.
creditrating = readtable("CreditRating_Historical.dat");
head(creditrating) ID WC_TA RE_TA EBIT_TA MVE_BVTD S_TA Industry Rating
_____ ______ ______ _______ ________ _____ ________ _______
62394 0.013 0.104 0.036 0.447 0.142 3 {'BB' }
48608 0.232 0.335 0.062 1.969 0.281 8 {'A' }
42444 0.311 0.367 0.074 1.935 0.366 1 {'A' }
48631 0.194 0.263 0.062 1.017 0.228 4 {'BBB'}
43768 0.121 0.413 0.057 3.647 0.466 12 {'AAA'}
39255 -0.117 -0.799 0.01 0.179 0.082 4 {'CCC'}
62236 0.087 0.158 0.049 0.816 0.324 2 {'BBB'}
39354 0.005 0.181 0.034 2.597 0.388 7 {'AA' }
Because each value in the ID variable is a unique customer ID, that is, length(unique(creditrating.ID)) is equal to the number of observations in creditrating, the ID variable is a poor predictor. Remove the ID variable from the table, and convert the Industry variable to a categorical variable.
creditrating = removevars(creditrating,"ID");
creditrating.Industry = categorical(creditrating.Industry);Partition the data into training and test sets. Use approximately 85% of the observations for the model selection and hyperparameter tuning process, and 15% of the observations to test the performance of the final model returned by fitcauto on new data. Use cvpartition to partition the data.
rng("default") % For reproducibility of the partition c = cvpartition(creditrating.Rating,"Holdout",0.15); trainingIndices = training(c); % Indices for the training set testIndices = test(c); % Indices for the test set creditTrain = creditrating(trainingIndices,:); creditTest = creditrating(testIndices,:);
Perform Feature Selection
Before passing the training data to fitcauto, find the important predictors by using the fscchi2 function. Visualize the predictor scores by using the bar function. Because some scores can be Inf, and bar discards Inf values, plot the finite scores first and then plot a finite representation of the Inf scores in a different color.
[idx,scores] = fscchi2(creditTrain,"Rating"); bar(scores(idx)) % Represents finite scores hold on veryImportant = isinf(scores); finiteMax = max(scores(~veryImportant)); bar(finiteMax*veryImportant(idx)) % Represents Inf scores hold off xticklabels(strrep(creditTrain.Properties.VariableNames(idx),"_","\_")) xtickangle(45) legend(["Finite Scores","Inf Scores"])

Notice that the Industry predictor has a low score corresponding to a p-value that is greater than 0.05, which indicates that Industry might not be an important feature. Remove the Industry feature from the training and test data sets.
creditTrain = removevars(creditTrain,'Industry'); creditTest = removevars(creditTest,'Industry');
Run fitcauto
Pass the training data to fitcauto. The function uses Bayesian optimization to select models and their hyperparameter values, and returns a trained model Mdl with the best expected performance. Specify to try all available learner types and run the optimization in parallel (requires Parallel Computing Toolbox™). Return a second output Results that contains the details of the Bayesian optimization.
Expect this process to take some time. By default, fitcauto provides a plot of the optimization and an iterative display of the optimization results. For more information on how to interpret these results, see Verbose Display.
options = struct("UseParallel",true); [Mdl,Results] = fitcauto(creditTrain,"Rating", ... "Learners","all","HyperparameterOptimizationOptions",options);
Starting parallel pool (parpool) using the 'Processes' profile ... 10-May-2024 11:22:29: Job Queued. Waiting for parallel pool job with ID 2 to start ... Connected to parallel pool with 6 workers. Copying objective function to workers... Done copying objective function to workers. Learner types to explore: discr, ensemble, kernel, knn, linear, nb, net, svm, tree Total iterations (MaxObjectiveEvaluations): 270 Total time (MaxTime): Inf |=======================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | validation loss | | | |=======================================================================================================================================================| | 1 | 6 | Best | 0.29704 | 3.5043 | 0.29704 | 0.29704 | tree | MinLeafSize: 9 | | 2 | 6 | Accept | 0.46844 | 3.6329 | 0.29704 | 0.29704 | knn | NumNeighbors: 2 | | | | | | | | | | Distance: cosine | | | | | | | | | | Standardize: false | | 3 | 6 | Accept | 0.47383 | 6.097 | 0.29704 | 0.29704 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.65536 | | | | | | | | | | KernelScale: 3.7007 | | | | | | | | | | Standardize: false | | 4 | 5 | Best | 0.26383 | 6.3788 | 0.26383 | 0.28023 | net | Activations: none | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.00069303 | | | | | | | | | | LayerSizes: 4 | | 5 | 5 | Accept | 0.26533 | 1.2671 | 0.26383 | 0.28023 | tree | MinLeafSize: 48 | | 6 | 5 | Accept | 0.26742 | 0.64789 | 0.26383 | 0.28023 | knn | NumNeighbors: 197 | | | | | | | | | | Distance: cityblock | | | | | | | | | | Standardize: false | | 7 | 5 | Accept | 0.32994 | 0.29613 | 0.26383 | 0.29963 | tree | MinLeafSize: 3 | | 8 | 6 | Accept | 0.68202 | 1.3718 | 0.26383 | 0.29963 | knn | NumNeighbors: 165 | | | | | | | | | | Distance: jaccard | | | | | | | | | | Standardize: false | | 9 | 6 | Accept | 0.68202 | 1.345 | 0.26383 | 0.29963 | knn | NumNeighbors: 165 | | | | | | | | | | Distance: jaccard | | | | | | | | | | Standardize: false | | 10 | 6 | Accept | 0.28059 | 0.79392 | 0.26383 | 0.28059 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 11 | 6 | Accept | 0.73886 | 6.3436 | 0.26383 | 0.28059 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 0.073664 | | | | | | | | | | Lambda: 0.008931 | | | | | | | | | | Standardize: true | | 12 | 6 | Accept | 0.74185 | 4.3986 | 0.26383 | 0.28059 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 0.0016013 | | | | | | | | | | Lambda: 0.043107 | | | | | | | | | | Standardize: false | | 13 | 6 | Accept | 0.28627 | 0.79497 | 0.26383 | 0.28059 | tree | MinLeafSize: 10 | | 14 | 6 | Best | 0.24529 | 2.6341 | 0.24529 | 0.24529 | linear | Coding: onevsone | | | | | | | | | | Lambda: 8.6988e-06 | | | | | | | | | | Learner: svm | | 15 | 6 | Best | 0.242 | 16.312 | 0.242 | 0.24529 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 1.705e-06 | | | | | | | | | | LayerSizes: 2 | | 16 | 5 | Accept | 0.3799 | 0.186 | 0.242 | 0.24529 | tree | MinLeafSize: 492 | | 17 | 5 | Accept | 0.27281 | 33.527 | 0.242 | 0.24529 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 3.3048e-07 | | | | | | | | | | LayerSizes: 34 | | 18 | 6 | Accept | 0.63865 | 0.39589 | 0.242 | 0.24529 | knn | NumNeighbors: 31 | | | | | | | | | | Distance: hamming | | | | | | | | | | Standardize: false | | 19 | 6 | Accept | 0.63865 | 1.7736 | 0.242 | 0.24529 | knn | NumNeighbors: 31 | | | | | | | | | | Distance: hamming | | | | | | | | | | Standardize: false | | 20 | 6 | Accept | 0.74185 | 1.0453 | 0.242 | 0.24529 | discr | Delta: 217.25 | | | | | | | | | | Gamma: 0.30834 | |=======================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | validation loss | | | |=======================================================================================================================================================| | 21 | 4 | Accept | 0.28627 | 29.738 | 0.242 | 0.24529 | ensemble | Coding: onevsone | | | | | | | | | | Method: GentleBoost | | | | | | | | | | NumLearningCycles: 36 | | | | | | | | | | LearnRate: 0.13026 | | | | | | | | | | MinLeafSize: 2 | | 22 | 4 | Accept | 0.24589 | 5.2013 | 0.242 | 0.24529 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 0.9881 | | | | | | | | | | Lambda: 3.9237e-05 | | | | | | | | | | Standardize: false | | 23 | 4 | Accept | 0.42746 | 0.15749 | 0.242 | 0.24529 | discr | Delta: 0.0016931 | | | | | | | | | | Gamma: 0.85944 | | 24 | 6 | Accept | 0.47742 | 0.94715 | 0.242 | 0.25846 | linear | Coding: onevsall | | | | | | | | | | Lambda: 2.4303e-06 | | | | | | | | | | Learner: svm | | 25 | 6 | Accept | 0.47742 | 1.748 | 0.242 | 0.25846 | linear | Coding: onevsall | | | | | | | | | | Lambda: 2.4303e-06 | | | | | | | | | | Learner: svm | | 26 | 5 | Accept | 0.47742 | 2.6179 | 0.242 | 0.28059 | linear | Coding: onevsall | | | | | | | | | | Lambda: 2.4303e-06 | | | | | | | | | | Learner: svm | | 27 | 5 | Accept | 0.74185 | 0.58088 | 0.242 | 0.28059 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.20318 | | | | | | | | | | LayerSizes: 4 | | 28 | 5 | Accept | 0.67424 | 5.7245 | 0.242 | 0.28059 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 4.9742 | | | | | | | | | | Lambda: 0.12271 | | | | | | | | | | Standardize: false | | 29 | 5 | Accept | 0.24379 | 2.0796 | 0.242 | 0.28059 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 0.18363 | | | | | | | | | | KernelScale: 0.068442 | | | | | | | | | | Standardize: false | | 30 | 6 | Accept | 0.56087 | 2.7403 | 0.242 | 0.28059 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.020179 | | | | | | | | | | KernelScale: 45.159 | | | | | | | | | | Standardize: true | | 31 | 6 | Accept | 0.24708 | 1.9624 | 0.242 | 0.28059 | linear | Coding: onevsone | | | | | | | | | | Lambda: 0.0025611 | | | | | | | | | | Learner: logistic | | 32 | 6 | Accept | 0.29704 | 0.20854 | 0.242 | 0.28059 | tree | MinLeafSize: 9 | | 33 | 6 | Accept | 0.74185 | 47.894 | 0.242 | 0.28059 | ensemble | Coding: onevsone | | | | | | | | | | Method: LogitBoost | | | | | | | | | | NumLearningCycles: 77 | | | | | | | | | | LearnRate: 0.0054224 | | | | | | | | | | MinLeafSize: 1430 | | 34 | 5 | Accept | 0.74185 | 4.6386 | 0.242 | 0.28059 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 0.0072048 | | | | | | | | | | Lambda: 0.028163 | | | | | | | | | | Standardize: false | | 35 | 5 | Accept | 0.55639 | 0.65802 | 0.242 | 0.28059 | discr | Delta: 1.0613 | | | | | | | | | | Gamma: 0.24003 | | 36 | 4 | Accept | 0.24379 | 7.4023 | 0.242 | 0.28059 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 1.7808 | | | | | | | | | | Lambda: 3.2594e-05 | | | | | | | | | | Standardize: false | | 37 | 4 | Accept | 0.24678 | 7.2124 | 0.242 | 0.28059 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 1.7808 | | | | | | | | | | Lambda: 3.2594e-05 | | | | | | | | | | Standardize: false | | 38 | 3 | Accept | 0.67155 | 1.7195 | 0.242 | 0.28059 | knn | NumNeighbors: 1445 | | | | | | | | | | Distance: spearman | | | | | | | | | | Standardize: true | | 39 | 3 | Accept | 0.46455 | 0.60315 | 0.242 | 0.28059 | linear | Coding: onevsall | | | | | | | | | | Lambda: 0.3312 | | | | | | | | | | Learner: logistic | | 40 | 6 | Accept | 0.28059 | 0.14968 | 0.242 | 0.28059 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | |=======================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | validation loss | | | |=======================================================================================================================================================| | 41 | 5 | Accept | 0.61113 | 0.31292 | 0.242 | 0.28059 | tree | MinLeafSize: 881 | | 42 | 5 | Accept | 0.28059 | 0.75648 | 0.242 | 0.28059 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 43 | 5 | Accept | 0.29494 | 1.4567 | 0.242 | 0.28059 | linear | Coding: onevsone | | | | | | | | | | Lambda: 0.03378 | | | | | | | | | | Learner: svm | | 44 | 5 | Accept | 0.43823 | 0.74458 | 0.242 | 0.28059 | linear | Coding: onevsall | | | | | | | | | | Lambda: 0.0084424 | | | | | | | | | | Learner: logistic | | 45 | 5 | Accept | 0.71702 | 0.49019 | 0.242 | 0.28059 | knn | NumNeighbors: 457 | | | | | | | | | | Distance: jaccard | | | | | | | | | | Standardize: true | | 46 | 6 | Accept | 0.24738 | 2.2676 | 0.242 | 0.28059 | linear | Coding: onevsone | | | | | | | | | | Lambda: 1.8074e-05 | | | | | | | | | | Learner: logistic | | 47 | 4 | Accept | 0.45648 | 8.8659 | 0.242 | 0.26483 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.0075021 | | | | | | | | | | KernelScale: 0.023323 | | | | | | | | | | Standardize: true | | 48 | 4 | Accept | 0.47203 | 4.1096 | 0.242 | 0.26483 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 1.4724 | | | | | | | | | | Lambda: 0.063523 | | | | | | | | | | Standardize: true | | 49 | 4 | Accept | 0.24738 | 3.0612 | 0.242 | 0.26483 | linear | Coding: onevsone | | | | | | | | | | Lambda: 1.8074e-05 | | | | | | | | | | Learner: logistic | | 50 | 4 | Accept | 0.24768 | 1.7307 | 0.242 | 0.26483 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 0.013982 | | | | | | | | | | KernelScale: 0.1266 | | | | | | | | | | Standardize: true | | 51 | 6 | Accept | 0.74185 | 2.788 | 0.242 | 0.26483 | nb | DistributionNames: kernel | | | | | | | | | | Width: 417.63 | | | | | | | | | | Standardize: false | | 52 | 6 | Accept | 0.42716 | 0.11455 | 0.242 | 0.26483 | discr | Delta: 0.021092 | | | | | | | | | | Gamma: 0.256 | | 53 | 6 | Accept | 0.24349 | 76.191 | 0.242 | 0.26483 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 1.3715e-05 | | | | | | | | | | LayerSizes: 122 | | 54 | 6 | Accept | 0.43314 | 0.9632 | 0.242 | 0.26483 | discr | Delta: 9.1157e-06 | | | | | | | | | | Gamma: 0.53332 | | 55 | 6 | Accept | 0.74185 | 4.3906 | 0.242 | 0.26483 | nb | DistributionNames: kernel | | | | | | | | | | Width: 18.623 | | | | | | | | | | Standardize: true | | 56 | 6 | Accept | 0.45917 | 8.4506 | 0.242 | 0.26483 | net | Activations: relu | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.007474 | | | | | | | | | | LayerSizes: [ 1 20 ] | | 57 | 6 | Accept | 0.25456 | 0.31225 | 0.242 | 0.26483 | knn | NumNeighbors: 21 | | | | | | | | | | Distance: euclidean | | | | | | | | | | Standardize: false | | 58 | 6 | Accept | 0.24319 | 20.176 | 0.242 | 0.26454 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.5101e-05 | | | | | | | | | | LayerSizes: 17 | | 59 | 6 | Accept | 0.38139 | 1.3557 | 0.242 | 0.26454 | knn | NumNeighbors: 112 | | | | | | | | | | Distance: cosine | | | | | | | | | | Standardize: true | | 60 | 6 | Accept | 0.48699 | 2.5673 | 0.242 | 0.26188 | linear | Coding: onevsall | | | | | | | | | | Lambda: 0.85779 | | | | | | | | | | Learner: svm | |=======================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | validation loss | | | |=======================================================================================================================================================| | 61 | 6 | Accept | 0.24439 | 1.8066 | 0.242 | 0.25682 | linear | Coding: onevsone | | | | | | | | | | Lambda: 1.7606e-08 | | | | | | | | | | Learner: svm | | 62 | 6 | Accept | 0.54442 | 1.1095 | 0.242 | 0.25682 | knn | NumNeighbors: 1256 | | | | | | | | | | Distance: cosine | | | | | | | | | | Standardize: true | | 63 | 6 | Accept | 0.28059 | 0.25665 | 0.242 | 0.25682 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 64 | 6 | Accept | 0.78283 | 9.2719 | 0.242 | 0.25682 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 0.0084622 | | | | | | | | | | Lambda: 4.3995e-05 | | | | | | | | | | Standardize: true | | 65 | 6 | Accept | 0.43284 | 9.9414 | 0.242 | 0.25682 | ensemble | Method: Bag | | | | | | | | | | NumLearningCycles: 171 | | | | | | | | | | LearnRate: NaN | | | | | | | | | | MinLeafSize: 583 | | 66 | 6 | Accept | 0.56985 | 0.16211 | 0.242 | 0.25682 | discr | Delta: 1.0473 | | | | | | | | | | Gamma: 0.72754 | | 67 | 6 | Accept | 0.24559 | 17.448 | 0.242 | 0.25682 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 0.30375 | | | | | | | | | | KernelScale: 0.094485 | | | | | | | | | | Standardize: true | | 68 | 6 | Accept | 0.46246 | 2.2288 | 0.242 | 0.25682 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 15.954 | | | | | | | | | | KernelScale: 3.1701 | | | | | | | | | | Standardize: false | | 69 | 6 | Accept | 0.74065 | 4.0667 | 0.242 | 0.25682 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 0.027004 | | | | | | | | | | Lambda: 0.010714 | | | | | | | | | | Standardize: false | | 70 | 6 | Accept | 0.3117 | 117.25 | 0.242 | 0.25682 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 5.5502e-07 | | | | | | | | | | LayerSizes: 141 | | 71 | 6 | Accept | 0.37571 | 0.32565 | 0.242 | 0.25682 | tree | MinLeafSize: 469 | | 72 | 6 | Accept | 0.28836 | 2.5757 | 0.242 | 0.25682 | nb | DistributionNames: kernel | | | | | | | | | | Width: 0.049199 | | | | | | | | | | Standardize: true | | 73 | 6 | Accept | 0.71941 | 0.57072 | 0.242 | 0.25682 | knn | NumNeighbors: 681 | | | | | | | | | | Distance: jaccard | | | | | | | | | | Standardize: true | | 74 | 6 | Accept | 0.54472 | 0.57641 | 0.242 | 0.25682 | discr | Delta: 0.89876 | | | | | | | | | | Gamma: 0.34215 | | 75 | 6 | Accept | 0.53814 | 0.95437 | 0.242 | 0.25682 | ensemble | Method: Bag | | | | | | | | | | NumLearningCycles: 16 | | | | | | | | | | LearnRate: NaN | | | | | | | | | | MinLeafSize: 1027 | | 76 | 6 | Accept | 0.5175 | 3.6894 | 0.242 | 0.25682 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.0022428 | | | | | | | | | | LayerSizes: [ 65 2 ] | | 77 | 6 | Accept | 0.28896 | 1.2373 | 0.242 | 0.25682 | nb | DistributionNames: kernel | | | | | | | | | | Width: 0.026834 | | | | | | | | | | Standardize: true | | 78 | 6 | Accept | 0.74005 | 2.4 | 0.242 | 0.25682 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 408.15 | | | | | | | | | | Lambda: 0.00054653 | | | | | | | | | | Standardize: false | | 79 | 6 | Accept | 0.71463 | 2.3243 | 0.242 | 0.25682 | nb | DistributionNames: kernel | | | | | | | | | | Width: 5.7434 | | | | | | | | | | Standardize: false | | 80 | 6 | Accept | 0.31768 | 1.4475 | 0.242 | 0.25281 | linear | Coding: onevsone | | | | | | | | | | Lambda: 0.14117 | | | | | | | | | | Learner: svm | |=======================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | validation loss | | | |=======================================================================================================================================================| | 81 | 6 | Accept | 0.42926 | 0.10631 | 0.242 | 0.25281 | discr | Delta: 0.0056255 | | | | | | | | | | Gamma: 0.29449 | | 82 | 6 | Accept | 0.74185 | 3.7651 | 0.242 | 0.25281 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 0.0021877 | | | | | | | | | | Lambda: 0.12375 | | | | | | | | | | Standardize: true | | 83 | 6 | Accept | 0.27879 | 196.9 | 0.242 | 0.25281 | ensemble | Coding: onevsone | | | | | | | | | | Method: GentleBoost | | | | | | | | | | NumLearningCycles: 306 | | | | | | | | | | LearnRate: 0.0028268 | | | | | | | | | | MinLeafSize: 5 | | 84 | 6 | Accept | 0.69518 | 0.10299 | 0.242 | 0.25281 | discr | Delta: 2.2186 | | | | | | | | | | Gamma: 0.48692 | | 85 | 6 | Accept | 0.46515 | 0.10863 | 0.242 | 0.25281 | tree | MinLeafSize: 666 | | 86 | 6 | Accept | 0.25845 | 11.362 | 0.242 | 0.25281 | ensemble | Method: AdaBoostM2 | | | | | | | | | | NumLearningCycles: 232 | | | | | | | | | | LearnRate: 0.057232 | | | | | | | | | | MinLeafSize: 2 | | 87 | 6 | Accept | 0.4819 | 0.23471 | 0.242 | 0.25281 | knn | NumNeighbors: 18 | | | | | | | | | | Distance: correlation | | | | | | | | | | Standardize: false | | 88 | 6 | Accept | 0.59497 | 0.64377 | 0.242 | 0.25281 | knn | NumNeighbors: 52 | | | | | | | | | | Distance: spearman | | | | | | | | | | Standardize: true | | 89 | 6 | Accept | 0.25965 | 7.2545 | 0.242 | 0.25281 | ensemble | Method: Bag | | | | | | | | | | NumLearningCycles: 66 | | | | | | | | | | LearnRate: NaN | | | | | | | | | | MinLeafSize: 1 | | 90 | 6 | Accept | 0.74185 | 1.158 | 0.242 | 0.25281 | net | Activations: tanh | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 8.4228 | | | | | | | | | | LayerSizes: [ 75 2 87 ] | | 91 | 6 | Accept | 0.26653 | 1.1482 | 0.242 | 0.25281 | ensemble | Method: AdaBoostM2 | | | | | | | | | | NumLearningCycles: 15 | | | | | | | | | | LearnRate: 0.11472 | | | | | | | | | | MinLeafSize: 39 | | 92 | 6 | Accept | 0.45049 | 5.8162 | 0.242 | 0.25281 | ensemble | Method: Bag | | | | | | | | | | NumLearningCycles: 127 | | | | | | | | | | LearnRate: NaN | | | | | | | | | | MinLeafSize: 830 | | 93 | 6 | Accept | 0.29225 | 5.0049 | 0.242 | 0.25281 | ensemble | Method: AdaBoostM2 | | | | | | | | | | NumLearningCycles: 110 | | | | | | | | | | LearnRate: 0.0018187 | | | | | | | | | | MinLeafSize: 15 | | 94 | 6 | Accept | 0.39097 | 0.88194 | 0.242 | 0.25281 | nb | DistributionNames: kernel | | | | | | | | | | Width: 0.0020466 | | | | | | | | | | Standardize: true | | 95 | 6 | Accept | 0.28059 | 0.16411 | 0.242 | 0.25281 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 96 | 6 | Accept | 0.74185 | 0.75488 | 0.242 | 0.25281 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.010761 | | | | | | | | | | LayerSizes: [ 18 8 17 ] | | 97 | 6 | Accept | 0.28059 | 0.11286 | 0.242 | 0.25281 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 98 | 6 | Accept | 0.28059 | 0.081407 | 0.242 | 0.25281 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 99 | 6 | Accept | 0.28627 | 0.21186 | 0.242 | 0.25281 | tree | MinLeafSize: 10 | | 100 | 6 | Accept | 0.74185 | 2.2671 | 0.242 | 0.25281 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 0.0013533 | | | | | | | | | | Lambda: 0.031901 | | | | | | | | | | Standardize: false | |=======================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | validation loss | | | |=======================================================================================================================================================| | 101 | 6 | Accept | 0.61322 | 1.0461 | 0.242 | 0.25281 | knn | NumNeighbors: 435 | | | | | | | | | | Distance: spearman | | | | | | | | | | Standardize: true | | 102 | 6 | Accept | 0.28059 | 0.12504 | 0.242 | 0.25281 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 103 | 6 | Accept | 0.28059 | 0.12053 | 0.242 | 0.25281 | nb | DistributionNames: normal | | | | | | | | | | Width: NaN | | | | | | | | | | Standardize: - | | 104 | 6 | Accept | 0.45169 | 0.81334 | 0.242 | 0.25215 | linear | Coding: onevsall | | | | | | | | | | Lambda: 0.017986 | | | | | | | | | | Learner: svm | | 105 | 6 | Accept | 0.54831 | 0.25986 | 0.242 | 0.25215 | knn | NumNeighbors: 50 | | | | | | | | | | Distance: correlation | | | | | | | | | | Standardize: true | | 106 | 6 | Accept | 0.4496 | 324.95 | 0.242 | 0.25215 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 0.0012647 | | | | | | | | | | KernelScale: 0.0017986 | | | | | | | | | | Standardize: true | | 107 | 6 | Accept | 0.3781 | 1.8041 | 0.242 | 0.25434 | linear | Coding: onevsall | | | | | | | | | | Lambda: 4.5023e-06 | | | | | | | | | | Learner: logistic | | 108 | 6 | Accept | 0.44451 | 0.70126 | 0.242 | 0.25462 | linear | Coding: onevsall | | | | | | | | | | Lambda: 0.12817 | | | | | | | | | | Learner: logistic | | 109 | 6 | Accept | 0.26025 | 23.436 | 0.242 | 0.25462 | ensemble | Coding: onevsall | | | | | | | | | | Method: AdaBoostM1 | | | | | | | | | | NumLearningCycles: 72 | | | | | | | | | | LearnRate: 0.015914 | | | | | | | | | | MinLeafSize: 134 | | 110 | 6 | Accept | 0.74185 | 0.55177 | 0.242 | 0.25462 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.16788 | | | | | | | | | | LayerSizes: [ 1 12 56 ] | | 111 | 6 | Accept | 0.4143 | 10.391 | 0.242 | 0.25462 | ensemble | Method: Bag | | | | | | | | | | NumLearningCycles: 175 | | | | | | | | | | LearnRate: NaN | | | | | | | | | | MinLeafSize: 563 | | 112 | 6 | Accept | 0.56506 | 0.11833 | 0.242 | 0.25462 | discr | Delta: 1.3228 | | | | | | | | | | Gamma: 0.34287 | | 113 | 6 | Accept | 0.64433 | 0.30407 | 0.242 | 0.25462 | knn | NumNeighbors: 26 | | | | | | | | | | Distance: hamming | | | | | | | | | | Standardize: false | | 114 | 6 | Accept | 0.27789 | 1.4143 | 0.242 | 0.25462 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 0.12591 | | | | | | | | | | KernelScale: 2.1296 | | | | | | | | | | Standardize: true | | 115 | 6 | Accept | 0.47113 | 0.16139 | 0.242 | 0.25462 | tree | MinLeafSize: 674 | | 116 | 6 | Accept | 0.74185 | 1.991 | 0.242 | 0.25462 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 525.99 | | | | | | | | | | Lambda: 0.024827 | | | | | | | | | | Standardize: true | | 117 | 6 | Accept | 0.45319 | 15.81 | 0.242 | 0.25462 | svm | Coding: onevsall | | | | | | | | | | BoxConstraint: 29.239 | | | | | | | | | | KernelScale: 0.37967 | | | | | | | | | | Standardize: false | | 118 | 6 | Accept | 0.47861 | 498.57 | 0.242 | 0.25462 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 127.44 | | | | | | | | | | KernelScale: 0.0098561 | | | | | | | | | | Standardize: true | | 119 | 6 | Accept | 0.32785 | 0.23811 | 0.242 | 0.25462 | tree | MinLeafSize: 4 | | 120 | 6 | Accept | 0.42985 | 0.1125 | 0.242 | 0.25462 | discr | Delta: 0.00023581 | | | | | | | | | | Gamma: 0.74169 | |=======================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | validation loss | | | |=======================================================================================================================================================| | 121 | 6 | Accept | 0.74185 | 0.28212 | 0.242 | 0.25462 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 2.7198 | | | | | | | | | | LayerSizes: 3 | | 122 | 6 | Accept | 0.29016 | 12.52 | 0.242 | 0.25462 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 0.24974 | | | | | | | | | | Lambda: 1.6898e-06 | | | | | | | | | | Standardize: false | | 123 | 6 | Accept | 0.61681 | 0.082854 | 0.242 | 0.25462 | tree | MinLeafSize: 1037 | | 124 | 6 | Accept | 0.28687 | 35.279 | 0.242 | 0.25462 | ensemble | Coding: onevsall | | | | | | | | | | Method: LogitBoost | | | | | | | | | | NumLearningCycles: 68 | | | | | | | | | | LearnRate: 0.99683 | | | | | | | | | | MinLeafSize: 9 | | 125 | 6 | Accept | 0.3793 | 0.16321 | 0.242 | 0.25462 | tree | MinLeafSize: 430 | | 126 | 6 | Accept | 0.2426 | 34.709 | 0.242 | 0.25462 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.0002e-07 | | | | | | | | | | LayerSizes: [ 2 128 ] | | 127 | 6 | Accept | 0.42656 | 0.12103 | 0.242 | 0.25462 | discr | Delta: 1.8328e-06 | | | | | | | | | | Gamma: 0.14399 | | 128 | 6 | Accept | 0.42537 | 0.089666 | 0.242 | 0.25462 | discr | Delta: 1.73e-05 | | | | | | | | | | Gamma: 0.15184 | | 129 | 6 | Accept | 0.24349 | 1.5715 | 0.242 | 0.25462 | svm | Coding: onevsone | | | | | | | | | | BoxConstraint: 6.9807 | | | | | | | | | | KernelScale: 0.60748 | | | | | | | | | | Standardize: false | | 130 | 6 | Accept | 0.79539 | 3.6791 | 0.242 | 0.25462 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 0.018788 | | | | | | | | | | Lambda: 1.0474e-05 | | | | | | | | | | Standardize: false | | 131 | 6 | Accept | 0.26862 | 11.273 | 0.242 | 0.25462 | ensemble | Method: AdaBoostM2 | | | | | | | | | | NumLearningCycles: 204 | | | | | | | | | | LearnRate: 0.0057787 | | | | | | | | | | MinLeafSize: 5 | | 132 | 6 | Accept | 0.48041 | 5.0926 | 0.242 | 0.25462 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 0.23411 | | | | | | | | | | Lambda: 3.608e-05 | | | | | | | | | | Standardize: true | | 133 | 6 | Accept | 0.30242 | 5.5709 | 0.242 | 0.25462 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 126.67 | | | | | | | | | | Lambda: 3.1052e-06 | | | | | | | | | | Standardize: false | | 134 | 6 | Accept | 0.24678 | 2.7052 | 0.242 | 0.25172 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.00059683 | | | | | | | | | | LayerSizes: 3 | | 135 | 6 | Accept | 0.73587 | 4.6448 | 0.242 | 0.25172 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 736.62 | | | | | | | | | | Lambda: 3.7849e-05 | | | | | | | | | | Standardize: true | | 136 | 6 | Accept | 0.4487 | 3.0531 | 0.242 | 0.25172 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 752.25 | | | | | | | | | | Lambda: 5.4668e-07 | | | | | | | | | | Standardize: false | | 137 | 6 | Best | 0.2414 | 11.239 | 0.2414 | 0.25462 | net | Activations: none | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.00046679 | | | | | | | | | | LayerSizes: [ 5 91 4 ] | | 138 | 6 | Accept | 0.25396 | 35.835 | 0.2414 | 0.25241 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 2.1901e-06 | | | | | | | | | | LayerSizes: [ 11 2 37 ] | | 139 | 6 | Accept | 0.45707 | 1.0704 | 0.2414 | 0.25462 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.00118 | | | | | | | | | | LayerSizes: [ 2 2 ] | | 140 | 6 | Accept | 0.2414 | 15.673 | 0.2414 | 0.24974 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 4.1041e-05 | | | | | | | | | | LayerSizes: [ 2 5 16 ] | |=======================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | validation loss | | | |=======================================================================================================================================================| | 141 | 6 | Accept | 0.24589 | 5.5905 | 0.2414 | 0.24974 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 6.9075 | | | | | | | | | | Lambda: 2.345e-05 | | | | | | | | | | Standardize: false | | 142 | 6 | Accept | 0.27311 | 6.419 | 0.2414 | 0.24974 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 1.1733 | | | | | | | | | | Lambda: 3.215e-05 | | | | | | | | | | Standardize: true | | 143 | 6 | Accept | 0.24649 | 5.5317 | 0.2414 | 0.24974 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 3.828 | | | | | | | | | | Lambda: 3.5924e-05 | | | | | | | | | | Standardize: false | | 144 | 6 | Accept | 0.42806 | 8.2774 | 0.2414 | 0.24974 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 0.093493 | | | | | | | | | | Lambda: 3.1039e-05 | | | | | | | | | | Standardize: false | | 145 | 6 | Accept | 0.37063 | 5.2086 | 0.2414 | 0.24974 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 102.4 | | | | | | | | | | Lambda: 3.5109e-05 | | | | | | | | | | Standardize: true | | 146 | 6 | Accept | 0.26234 | 5.3734 | 0.2414 | 0.24974 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 17.801 | | | | | | | | | | Lambda: 3.648e-05 | | | | | | | | | | Standardize: false | | 147 | 6 | Accept | 0.24888 | 5.7508 | 0.2414 | 0.24974 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 2.1707 | | | | | | | | | | Lambda: 1.0135e-05 | | | | | | | | | | Standardize: false | | 148 | 6 | Accept | 0.29674 | 4.1202 | 0.2414 | 0.24974 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 6.3863 | | | | | | | | | | Lambda: 3.9393e-05 | | | | | | | | | | Standardize: false | | 149 | 6 | Accept | 0.26413 | 7.9572 | 0.2414 | 0.24974 | ensemble | Coding: onevsall | | | | | | | | | | Method: LogitBoost | | | | | | | | | | NumLearningCycles: 22 | | | | | | | | | | LearnRate: 0.052662 | | | | | | | | | | MinLeafSize: 82 | | 150 | 6 | Accept | 0.29734 | 4.2389 | 0.2414 | 0.24974 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 6.1493 | | | | | | | | | | Lambda: 3.7581e-05 | | | | | | | | | | Standardize: false | | 151 | 6 | Accept | 0.29614 | 2.433 | 0.2414 | 0.24974 | ensemble | Method: AdaBoostM2 | | | | | | | | | | NumLearningCycles: 45 | | | | | | | | | | LearnRate: 0.037037 | | | | | | | | | | MinLeafSize: 215 | | 152 | 6 | Accept | 0.25187 | 6.677 | 0.2414 | 0.24974 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 4.4623 | | | | | | | | | | Lambda: 3.9302e-05 | | | | | | | | | | Standardize: true | | 153 | 6 | Accept | 0.24499 | 5.6011 | 0.2414 | 0.24974 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 3.6605 | | | | | | | | | | Lambda: 4.0695e-05 | | | | | | | | | | Standardize: false | | 154 | 6 | Best | 0.2399 | 10.914 | 0.2399 | 0.24458 | net | Activations: none | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.00087823 | | | | | | | | | | LayerSizes: [ 3 10 135 ] | | 155 | 6 | Accept | 0.26533 | 4.3657 | 0.2399 | 0.24458 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 3.1699 | | | | | | | | | | Lambda: 4.2302e-05 | | | | | | | | | | Standardize: false | | 156 | 6 | Accept | 0.25037 | 4.4619 | 0.2399 | 0.24458 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 1.4452 | | | | | | | | | | Lambda: 4.3079e-05 | | | | | | | | | | Standardize: false | | 157 | 6 | Accept | 0.24499 | 5.4242 | 0.2399 | 0.24458 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 3.7694 | | | | | | | | | | Lambda: 4.3534e-05 | | | | | | | | | | Standardize: false | | 158 | 6 | Accept | 0.26563 | 4.2206 | 0.2399 | 0.24458 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 3.1282 | | | | | | | | | | Lambda: 4.3446e-05 | | | | | | | | | | Standardize: false | | 159 | 6 | Accept | 0.24499 | 5.0307 | 0.2399 | 0.24458 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 4.0207 | | | | | | | | | | Lambda: 7.2479e-05 | | | | | | | | | | Standardize: false | | 160 | 6 | Accept | 0.24379 | 5.4303 | 0.2399 | 0.24458 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 4.6559 | | | | | | | | | | Lambda: 4.0574e-05 | | | | | | | | | | Standardize: false | |=======================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | validation loss | | | |=======================================================================================================================================================| | 161 | 6 | Accept | 0.24738 | 4.2359 | 0.2399 | 0.24458 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 1.8848 | | | | | | | | | | Lambda: 4.73e-05 | | | | | | | | | | Standardize: false | | 162 | 6 | Accept | 0.24738 | 5.267 | 0.2399 | 0.24458 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 0.80973 | | | | | | | | | | Lambda: 0.00016926 | | | | | | | | | | Standardize: false | | 163 | 6 | Accept | 0.24858 | 3.8912 | 0.2399 | 0.24458 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 2.002 | | | | | | | | | | Lambda: 6.9602e-05 | | | | | | | | | | Standardize: false | | 164 | 6 | Accept | 0.25396 | 3.3102 | 0.2399 | 0.24458 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 1.7161 | | | | | | | | | | Lambda: 0.00024265 | | | | | | | | | | Standardize: false | | 165 | 6 | Accept | 0.32486 | 3.3696 | 0.2399 | 0.24458 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 5.1868 | | | | | | | | | | Lambda: 0.00030745 | | | | | | | | | | Standardize: false | | 166 | 6 | Accept | 0.26593 | 6.9799 | 0.2399 | 0.24458 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 1.8249 | | | | | | | | | | Lambda: 5.4968e-05 | | | | | | | | | | Standardize: true | | 167 | 6 | Accept | 0.24858 | 4.795 | 0.2399 | 0.24458 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 1.0636 | | | | | | | | | | Lambda: 0.00040238 | | | | | | | | | | Standardize: false | | 168 | 6 | Accept | 0.25755 | 6.1694 | 0.2399 | 0.24458 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 2.245 | | | | | | | | | | Lambda: 6.6618e-05 | | | | | | | | | | Standardize: true | | 169 | 6 | Accept | 0.26234 | 3.0205 | 0.2399 | 0.24458 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 1.6204 | | | | | | | | | | Lambda: 0.00044561 | | | | | | | | | | Standardize: false | | 170 | 6 | Accept | 0.27161 | 3.0247 | 0.2399 | 0.24458 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 2.0168 | | | | | | | | | | Lambda: 0.00041694 | | | | | | | | | | Standardize: false | | 171 | 6 | Accept | 0.25217 | 8.6916 | 0.2399 | 0.24458 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 3.9761 | | | | | | | | | | Lambda: 8.9879e-06 | | | | | | | | | | Standardize: true | | 172 | 6 | Accept | 0.26653 | 4.4537 | 0.2399 | 0.24458 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 3.5929 | | | | | | | | | | Lambda: 0.00096967 | | | | | | | | | | Standardize: false | | 173 | 6 | Accept | 0.25666 | 6.5672 | 0.2399 | 0.24458 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 0.40713 | | | | | | | | | | Lambda: 4.6217e-05 | | | | | | | | | | Standardize: false | | 174 | 6 | Accept | 0.33204 | 3.0506 | 0.2399 | 0.24458 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 4.4378 | | | | | | | | | | Lambda: 0.00050652 | | | | | | | | | | Standardize: false | | 175 | 6 | Accept | 0.28866 | 2.9742 | 0.2399 | 0.24458 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 2.3933 | | | | | | | | | | Lambda: 0.00058278 | | | | | | | | | | Standardize: false | | 176 | 6 | Accept | 0.25217 | 4.854 | 0.2399 | 0.24458 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 4.8744 | | | | | | | | | | Lambda: 0.00030213 | | | | | | | | | | Standardize: false | | 177 | 6 | Accept | 0.37212 | 2.4693 | 0.2399 | 0.24458 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 3.8515 | | | | | | | | | | Lambda: 0.0029253 | | | | | | | | | | Standardize: false | | 178 | 6 | Accept | 0.24888 | 2.9753 | 0.2399 | 0.24458 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 1.0426 | | | | | | | | | | Lambda: 0.00064459 | | | | | | | | | | Standardize: false | | 179 | 6 | Accept | 0.25695 | 5.1847 | 0.2399 | 0.24458 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 4.8233 | | | | | | | | | | Lambda: 0.00023726 | | | | | | | | | | Standardize: true | | 180 | 6 | Accept | 0.26324 | 4.8044 | 0.2399 | 0.24458 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 4.8311 | | | | | | | | | | Lambda: 0.00054379 | | | | | | | | | | Standardize: true | |=======================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | validation loss | | | |=======================================================================================================================================================| | 181 | 6 | Accept | 0.24708 | 4.3993 | 0.2399 | 0.24458 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 0.97694 | | | | | | | | | | Lambda: 0.0030881 | | | | | | | | | | Standardize: false | | 182 | 6 | Accept | 0.25337 | 2.4427 | 0.2399 | 0.24714 | net | Activations: none | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.001292 | | | | | | | | | | LayerSizes: [ 26 4 ] | | 183 | 6 | Accept | 0.39067 | 2.5062 | 0.2399 | 0.24634 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 1.2819 | | | | | | | | | | Lambda: 0.0052145 | | | | | | | | | | Standardize: true | | 184 | 6 | Accept | 0.24678 | 4.7713 | 0.2399 | 0.24774 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 0.87503 | | | | | | | | | | Lambda: 0.0005457 | | | | | | | | | | Standardize: false | | 185 | 6 | Accept | 0.32725 | 3.0177 | 0.2399 | 0.24622 | kernel | Coding: onevsall | | | | | | | | | | KernelScale: 1.1776 | | | | | | | | | | Lambda: 0.00077561 | | | | | | | | | | Standardize: true | | 186 | 6 | Accept | 0.25486 | 5.2826 | 0.2399 | 0.24654 | kernel | Coding: onevsone | | | | | | | | | | KernelScale: 5.1429 | | | | | | | | | | Lambda: 0.00014913 | | | | | | | | | | Standardize: true | | 187 | 6 | Accept | 0.2405 | 13.417 | 0.2399 | 0.24374 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.00098014 | | | | | | | | | | LayerSizes: [ 3 168 6 ] | | 188 | 6 | Accept | 0.26772 | 5.7891 | 0.2399 | 0.24613 | net | Activations: none | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.0016998 | | | | | | | | | | LayerSizes: [ 1 222 ] | | 189 | 6 | Accept | 0.242 | 28.819 | 0.2399 | 0.24346 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.0765e-05 | | | | | | | | | | LayerSizes: [ 2 34 ] | | 190 | 6 | Accept | 0.2399 | 17.431 | 0.2399 | 0.2443 | net | Activations: relu | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.799e-05 | | | | | | | | | | LayerSizes: [ 6 2 13 ] | | 191 | 6 | Accept | 0.64373 | 4.0354 | 0.2399 | 0.24597 | net | Activations: relu | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 9.6616e-06 | | | | | | | | | | LayerSizes: [ 1 1 16 ] | | 192 | 6 | Accept | 0.29195 | 7.1387 | 0.2399 | 0.24504 | net | Activations: none | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.0018314 | | | | | | | | | | LayerSizes: [ 279 1 1 ] | | 193 | 6 | Accept | 0.24738 | 17.111 | 0.2399 | 0.24654 | net | Activations: relu | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 2.7262e-05 | | | | | | | | | | LayerSizes: [ 2 3 12 ] | | 194 | 6 | Accept | 0.25695 | 19.097 | 0.2399 | 0.2451 | net | Activations: tanh | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.0018592 | | | | | | | | | | LayerSizes: [ 5 8 21 ] | | 195 | 6 | Accept | 0.30482 | 7.588 | 0.2399 | 0.24529 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 1.6205e-05 | | | | | | | | | | LayerSizes: [ 3 1 ] | | 196 | 6 | Accept | 0.2426 | 16.05 | 0.2399 | 0.24348 | net | Activations: none | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 4.2692e-05 | | | | | | | | | | LayerSizes: [ 1 61 13 ] | | 197 | 6 | Accept | 0.25606 | 37.701 | 0.2399 | 0.24654 | net | Activations: relu | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 3.4757e-06 | | | | | | | | | | LayerSizes: [ 6 90 4 ] | | 198 | 6 | Accept | 0.2405 | 44.824 | 0.2399 | 0.2427 | net | Activations: relu | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.00071975 | | | | | | | | | | LayerSizes: [ 2 14 102 ] | | 199 | 6 | Accept | 0.26533 | 5.2454 | 0.2399 | 0.24562 | net | Activations: none | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.0031705 | | | | | | | | | | LayerSizes: [ 6 17 145 ] | | 200 | 6 | Accept | 0.28687 | 2.6155 | 0.2399 | 0.24499 | net | Activations: none | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.004398 | | | | | | | | | | LayerSizes: [ 2 69 9 ] | |=======================================================================================================================================================| | Iter | Active | Eval | Validation | Time for training | Observed min | Estimated min | Learner | Hyperparameter: Value | | | workers | result | loss | & validation (sec)| validation loss | validation loss | | | |=======================================================================================================================================================| | 201 | 6 | Accept | 0.24738 | 18.077 | 0.2399 | 0.24507 | net | Activations: relu | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 1.5557e-05 | | | | | | | | | | LayerSizes: [ 3 12 13 ] | | 202 | 6 | Accept | 0.2402 | 10.51 | 0.2399 | 0.24258 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 4.2267e-05 | | | | | | | | | | LayerSizes: [ 1 7 5 ] | | 203 | 6 | Accept | 0.33712 | 16.279 | 0.2399 | 0.24512 | net | Activations: none | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.0066296 | | | | | | | | | | LayerSizes: [ 3 225 228 ] | | 204 | 6 | Accept | 0.74185 | 0.57522 | 0.2399 | 0.24528 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.0022415 | | | | | | | | | | LayerSizes: [ 4 5 2 ] | | 205 | 6 | Accept | 0.2411 | 9.1254 | 0.2399 | 0.24401 | net | Activations: none | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 3.6813e-09 | | | | | | | | | | LayerSizes: [ 1 9 16 ] | | 206 | 6 | Accept | 0.74185 | 1.6902 | 0.2399 | 0.24501 | net | Activations: sigmoid | | | | | | | | | | Standardize: true | | | | | | | | | | Lambda: 0.0018843 | | | | | | | | | | LayerSizes: [ 26 5 52 ] | | 207 | 6 | Accept | 0.74185 | 0.96211 | 0.2399 | 0.24422 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.0010998 | | | | | | | | | | LayerSizes: [ 24 4 6 ] | | 208 | 6 | Accept | 0.24559 | 20.386 | 0.2399 | 0.24513 | net | Activations: tanh | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 3.1073e-05 | | | | | | | | | | LayerSizes: [ 2 2 2 ] | | 209 | 6 | Accept | 0.28418 | 16.286 | 0.2399 | 0.24432 | net | Activations: tanh | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.00041184 | | | | | | | | | | LayerSizes: [ 3 5 2 ] | | 210 | 6 | Accept | 0.29494 | 21.129 | 0.2399 | 0.24208 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | | | | | Lambda: 0.00051161 | | | | | | | | | | LayerSizes: [ 11 29 20 ] | | 211 | 6 | Accept | 0.2411 | 68.677 | 0.2399 | 0.24394 | net | Activations: sigmoid | | | | | | | | | | Standardize: false | | | | | | ...
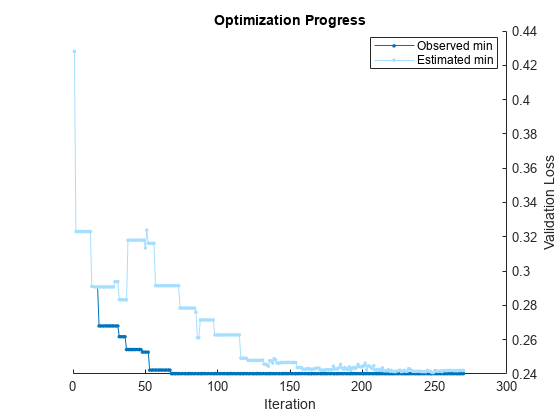
__________________________________________________________ Optimization completed. Total iterations: 270 Total elapsed time: 2056.3088 seconds Total time for training and validation: 11269.3988 seconds Best observed learner is a net model with: Learner: net Activations: none Standardize: true Lambda: 0.00080617 LayerSizes: [24 293 67] Observed validation loss: 0.23931 Time for training and validation: 35.4452 seconds Best estimated learner (returned model) is a net model with: Learner: net Activations: none Standardize: false Lambda: 0.001083 LayerSizes: [4 18 18] Estimated validation loss: 0.24198 Estimated time for training and validation: 6.818 seconds Documentation for fitcauto display
The final model returned by fitcauto corresponds to the best estimated learner. Before returning the model, the function retrains it using the entire training data (creditTrain), the listed Learner (or model) type, and the displayed hyperparameter values.
Evaluate Test Set Performance
The model Mdl corresponds to the best point in the Bayesian optimization according to the "min-visited-mean" criterion. To gauge how the model will perform on new data, look at the observed cross-validation accuracy of the model (cvAccuracy) and its general estimated performance based on the Bayesian optimization (estimatedAccuracy).
[x,~,iteration] = bestPoint(Results,"Criterion","min-visited-mean"); cvError = Results.ObjectiveTrace(iteration); cvAccuracy = 1 - cvError
cvAccuracy = 0.7604
estimatedError = predictObjective(Results,x); estimatedAccuracy = 1 - estimatedError
estimatedAccuracy = 0.7580
Evaluate the performance of the model on the test set. Create a confusion matrix from the results, and specify the order of the classes in the confusion matrix.
testAccuracy = 1 - loss(Mdl,creditTest,"Rating")testAccuracy = 0.7471
cm = confusionchart(creditTest.Rating,predict(Mdl,creditTest)); sortClasses(cm,["AAA","AA","A","BBB","BB","B","CCC"])
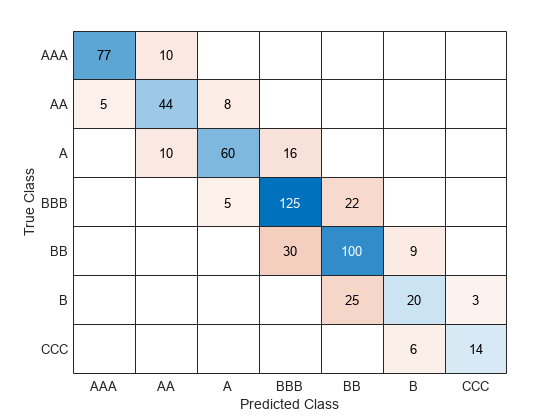
Input Arguments
Sample data, specified as a table. Each row of Tbl corresponds to one observation, and each column corresponds to one predictor. Optionally, Tbl can contain one additional column for the response variable. Multicolumn variables and cell arrays other than cell arrays of character vectors are not accepted.
If Tbl contains the response variable, and you want to use all remaining
variables in Tbl as predictors, specify the response variable using
ResponseVarName.
If Tbl contains the response variable, and you want to use only a subset of the remaining variables in Tbl as predictors, specify a formula using formula.
If Tbl does not contain the response variable, specify a response variable using Y. The length of the response variable and the number of rows in Tbl must be equal.
Data Types: table
Response variable name, specified as the name of a variable in
Tbl.
You must specify ResponseVarName as a character vector or string scalar.
For example, if the response variable Y is
stored as Tbl.Y, then specify it as
"Y". Otherwise, the software
treats all columns of Tbl, including
Y, as predictors when training
the model.
The response variable must be a categorical, character, or string array; a logical or numeric
vector; or a cell array of character vectors. If
Y is a character array, then each
element of the response variable must correspond to one row of
the array.
A good practice is to specify the order of the classes by using the
ClassNames name-value
argument.
Data Types: char | string
Explanatory model of the response variable and a subset of the predictor variables,
specified as a character vector or string scalar in the form
"Y~x1+x2+x3". In this form, Y represents the
response variable, and x1, x2, and
x3 represent the predictor variables.
To specify a subset of variables in Tbl as predictors for
training the model, use a formula. If you specify a formula, then the software does not
use any variables in Tbl that do not appear in
formula.
The variable names in the formula must be both variable names in Tbl
(Tbl.Properties.VariableNames) and valid MATLAB® identifiers. You can verify the variable names in Tbl by
using the isvarname function. If the variable names
are not valid, then you can convert them by using the matlab.lang.makeValidName function.
Data Types: char | string
Class labels, specified as a numeric, categorical, or logical vector, a character or string array, or a cell array of character vectors.
If
Yis a character array, then each element of the class labels must correspond to one row of the array.The length of
Ymust be equal to the number of rows inTblorX.A good practice is to specify the class order by using the
ClassNamesname-value argument.
Data Types: single | double | categorical | logical | char | string | cell
Predictor data, specified as a numeric matrix.
Each row of X corresponds to one observation, and each column corresponds to one predictor.
The length of Y and the number of rows in X must be equal.
To specify the names of the predictors in the order of their appearance in
X, use the PredictorNames name-value
argument.
Data Types: single | double
Note
The software treats NaN, empty character vector
(''), empty string (""),
<missing>, and <undefined> elements as
missing data. The software removes rows of data corresponding to missing values in the
response variable. However, the treatment of missing values in the predictor data
X or Tbl varies among models (or
learners).
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: "HyperparameterOptimizationOptions",struct("MaxObjectiveEvaluations",200,"Verbose",2)
specifies to run 200 iterations of the optimization process (that is, try 200 model
hyperparameter combinations), and to display information in the Command Window about the
next model hyperparameter combination to be evaluated.
Optimization Options
Types of classification models to try during the optimization, specified as a value in the first table below or one or more learner names in the second table. Specify multiple learner names as a string or cell array.
| Value | Description |
|---|---|
"auto" |
Note To provide the best hyperparameter optimization experience, the automatic selection of learners behavior is subject to frequent changes. For a more consistent selection of learners across software releases, explicitly specify the models you want to include. |
"all" | fitcauto selects all possible learners. |
"all-linear" | fitcauto selects linear learners:
"discr" (with a linear discriminant type) and
"linear". |
"all-nonlinear" | fitcauto selects all nonlinear learners:
"discr" (with a quadratic discriminant type),
"ensemble", "kernel",
"knn", "nb",
"net", "svm" (with a Gaussian or
polynomial kernel), and "tree". |
Note
For greater efficiency, fitcauto does not select the following combinations of models when you specify one of the previous values.
"kernel"and"svm"(with a Gaussian kernel) —fitcautochooses the first when the predictor data has more than 11,000 observations, and the second otherwise."linear"and"svm"(with a linear kernel) —fitcautochooses the first.
| Learner Name | Description |
|---|---|
"discr" | Discriminant analysis classifier |
"ensemble" | Ensemble classification model |
"kernel" | Kernel classification model |
"knn" | k-nearest neighbor model |
"linear" | Linear classification model |
"nb" | Naive Bayes classifier |
"net" | Neural network classifier |
"svm" | Support vector machine classifier |
"tree" | Binary decision classification tree |
Example: "Learners","all"
Example: "Learners","ensemble"
Example: "Learners",["svm","tree"]
Data Types: char | string | cell
Hyperparameters to optimize, specified as "auto" or
"all". The optimizable hyperparameters depend on the model (or
learner), as described in this table.
| Learner Name | Hyperparameters for "auto" | Additional Hyperparameters for "all" | Notes |
|---|---|---|---|
"discr" | Delta, Gamma | DiscrimType |
For more information, including hyperparameter search
ranges, see |
"ensemble" | Method, NumLearningCycles, LearnRate, MinLeafSize,
Coding (for ECOC
classification with three or more classes only) | MaxNumSplits,
NumVariablesToSample, SplitCriterion |
For more information, including hyperparameter search
ranges, see |
"kernel" | KernelScale, Lambda, Standardize, Coding (for three
or more classes only) | Learner, NumExpansionDimensions | For more information, including hyperparameter search ranges, see
OptimizeHyperparameters and OptimizeHyperparameters (for three or more classes only). Note
that you cannot change hyperparameter search ranges when you use
fitcauto. |
"knn" | Distance,
NumNeighbors,
Standardize | DistanceWeight, Exponent | For more information, including hyperparameter search ranges, see
OptimizeHyperparameters. Note that you cannot change
hyperparameter search ranges when you use
fitcauto. |
"linear" | Lambda, Learner, Coding (for three
or more classes only) | Regularization | For more information, including hyperparameter search ranges, see
OptimizeHyperparameters and OptimizeHyperparameters (for three or more classes only). Note
that you cannot change hyperparameter search ranges when you use
fitcauto. |
"nb" | DistributionNames, Standardize, Width | Kernel | For more information, including hyperparameter search ranges, see
OptimizeHyperparameters. Note that you cannot change
hyperparameter search ranges when you use
fitcauto. |
"net" | Activations, Lambda, LayerSizes, Standardize | LayerBiasesInitializer, LayerWeightsInitializer | For more information, including hyperparameter search ranges, see
OptimizeHyperparameters. Note that you cannot change
hyperparameter search ranges when you use
fitcauto. |
"svm" | BoxConstraint, KernelScale,
Standardize,
Coding (for three
or more classes only) | KernelFunction, PolynomialOrder | When the For more information, including hyperparameter search
ranges, see |
"tree" | MinLeafSize | MaxNumSplits,
SplitCriterion | For more information, including hyperparameter search ranges, see
OptimizeHyperparameters. Note that you cannot change
hyperparameter search ranges when you use
fitcauto. |
Note
When Learners is set to a value other than
"auto", the default values for the model hyperparameters not
being optimized match the default fit function values, unless otherwise indicated in
the table notes. When Learners is set to
"auto", the optimized hyperparameter search ranges and
nonoptimized hyperparameter values can vary, depending on the characteristics of the
training data. For more information, see Automatic Selection of Learners.
Example: "OptimizeHyperparameters","all"
Options for optimization, specified as a HyperparameterOptimizationOptions object or a structure. All the options
are optional. The options that you can set in a structure are the same as those in the
HyperparameterOptimizationOptions object.
| Option | Values | Default |
|---|---|---|
Optimizer |
| "bayesopt" |
ConstraintBounds | Constraint bounds for N optimization problems,
specified as an N-by-2 numeric matrix or
| [] |
ConstraintTarget | Constraint target for the optimization problems, specified as
| If you specify ConstraintBounds and
ConstraintType, then the default value is
"matlab". Otherwise, the default value is
[]. |
ConstraintType | Constraint type for the optimization problems, specified as
| [] |
LossFun | Type of validation loss to optimize, specified as
| "auto" |
MaxObjectiveEvaluations | Maximum number of objective function evaluations. If you specify multiple
optimization problems using ConstraintBounds, the value
of MaxObjectiveEvaluations applies to each optimization
problem individually. |
|
MaxTime | Time limit for the optimization, specified as a nonnegative real
scalar. The time limit is in seconds, as measured by | Inf |
ShowPlots | Logical value indicating whether to show plots of the optimization
progress. If this option is true, the software plots the
observed minimum validation loss against the iteration number. When you use
Bayesian optimization, the plot also shows the estimated minimum validation
loss. | true |
SaveIntermediateResults | Logical value indicating whether to save the optimization results. If
this option is true, the software overwrites a workspace
variable at each iteration. The variable is a SupervisedLearningBayesianOptimization object named
SupervisedLearningBayesoptResults if you use Bayesian
optimization, and a table named ASHAResults if you use ASHA
optimization. If you specify multiple optimization problems using
ConstraintBounds, the workspace variable is an AggregateBayesianOptimization object named
AggregateBayesoptResults. | false |
Verbose | Display level at the command line:
| 1 |
UseParallel | Logical value indicating whether to run the optimization in parallel, which requires Parallel Computing Toolbox™. Due to the nonreproducibility of parallel timing, parallel optimization does not necessarily yield reproducible results. For details, see Parallel Bayesian Optimization. | false |
Repartition | Logical value indicating whether to repartition the
cross-validation at every iteration. If this option is
A value of | false |
MaxTrainingSetSize | Maximum number of observations in each training set, specified as a positive integer. This value matches the largest training set size. Note If you want to specify this value, the | Largest available training partition size
|
MinTrainingSetSize | Minimum number of observations in each training set, specified as a positive integer. This value is the lower bound for the smallest training set size. Note If you want to specify this value, the | 100 |
| Specify only one of the following three options. | ||
CVPartition | cvpartition object created by cvpartition | Kfold=5 if you do not specify a
cross-validation option |
Holdout | Scalar in the range (0,1) representing the holdout
fraction | |
Kfold | Integer greater than 1 | |
Example: HyperparameterOptimizationOptions=struct(UseParallel=true)
Classification Options
Categorical predictors list, specified as one of the values in this table.
| Value | Description |
|---|---|
| Vector of positive integers |
Each entry in the vector is an index value indicating that the corresponding predictor is
categorical. The index values are between 1 and If |
| Logical vector |
A |
| Character matrix | Each row of the matrix is the name of a predictor variable. The names must match the entries in PredictorNames. Pad the names with extra blanks so each row of the character matrix has the same length. |
| String array or cell array of character vectors | Each element in the array is the name of a predictor variable. The names must match the entries in PredictorNames. |
"all" | All predictors are categorical. |
By default, if the predictor data is in a table (Tbl),
fitcauto assumes that a variable is categorical if it is a
logical vector, categorical vector, character array, string array, or cell array of
character vectors. However, learners that use decision trees assume that mathematically
ordered categorical vectors are continuous variables. If the predictor data is a matrix
(X), fitcauto assumes that all
predictors are continuous. To identify any other predictors as categorical predictors,
specify them by using the CategoricalPredictors name-value
argument.
For more information on how fitting functions treat categorical predictors, see Automatic Creation of Dummy Variables.
Note
fitcautodoes not support categorical predictors for discriminant analysis classifiers. That is, if you wantLearnersto include"discr"models, you cannot specify theCategoricalPredictorsname-value argument or use a table of sample data (Tbl) containing categorical predictors.fitcautodoes not support a mix of numeric and categorical predictors for k-nearest neighbor models. That is, if you wantLearnersto include"knn"models, you must specify theCategoricalPredictorsvalue as"all"or[].
Example: "CategoricalPredictors","all"
Data Types: single | double | logical | char | string | cell
Names of classes to use for training, specified as a categorical, character, or string
array; a logical or numeric vector; or a cell array of character vectors.
ClassNames must have the same data type as the response variable
in Tbl or Y.
If ClassNames is a character array, then each element must correspond to one row of the array.
Use ClassNames to:
Specify the order of the classes during training.
Specify the order of any input or output argument dimension that corresponds to the class order. For example, use
ClassNamesto specify the order of the dimensions ofCostor the column order of classification scores returned bypredict.Select a subset of classes for training. For example, suppose that the set of all distinct class names in
Yis["a","b","c"]. To train the model using observations from classes"a"and"c"only, specifyClassNames=["a","c"].
The default value for ClassNames is the set of all distinct class names in the response variable in Tbl or Y.
Example: ClassNames=["b","g"]
Data Types: categorical | char | string | logical | single | double | cell
Misclassification cost, specified as a square matrix or structure array.
If you specify a square matrix
Costand the true class of an observation isi, thenCost(i,j)is the cost of classifying a point into classj. That is, rows correspond to the true classes and columns correspond to the predicted classes. To specify the class order for the corresponding rows and columns ofCost, also specify theClassNamesname-value argument.If you specify a structure
S, then it must have two fields:S.ClassNames, which contains the class names as a variable of the same data type asYS.ClassificationCosts, which contains the cost matrix with rows and columns ordered as inS.ClassNames
In general, misclassification costs are used differently by the various models in
Learners. However, when you specify Cost
with the default LossFun value in the
HyperparameterOptimizationOptions object or structure, the
fitcauto function computes the same observed misclassification
cost ("classifcost") to compare the models during the optimization
process.
The default value for Cost is ones(K) –
eye(K), where K is the number of distinct
classes.
Example: "Cost",[0 1; 2 0]
Data Types: single | double | struct
Predictor variable names, specified as a string array of unique names or cell array of unique
character vectors. The functionality of PredictorNames depends on the
way you supply the training data.
If you supply
XandY, then you can usePredictorNamesto assign names to the predictor variables inX.The order of the names in
PredictorNamesmust correspond to the column order ofX. That is,PredictorNames{1}is the name ofX(:,1),PredictorNames{2}is the name ofX(:,2), and so on. Also,size(X,2)andnumel(PredictorNames)must be equal.By default,
PredictorNamesis{'x1','x2',...}.
If you supply
Tbl, then you can usePredictorNamesto choose which predictor variables to use in training. That is,fitcautouses only the predictor variables inPredictorNamesand the response variable during training.PredictorNamesmust be a subset ofTbl.Properties.VariableNamesand cannot include the name of the response variable.By default,
PredictorNamescontains the names of all predictor variables.A good practice is to specify the predictors for training using either
PredictorNamesorformula, but not both.
Example: PredictorNames=["SepalLength","SepalWidth","PetalLength","PetalWidth"]
Data Types: string | cell
Prior probabilities for each class, specified as a value in this table.
| Value | Description |
|---|---|
"empirical" | The class prior probabilities are the class relative frequencies in
Y. |
"uniform" | All class prior probabilities are equal to 1/K, where K is the number of classes. |
| numeric vector | Each element is a class prior probability. Order the elements according
to Mdl.ClassNames or specify the
order using the ClassNames name-value argument. The
software normalizes the elements to sum to 1. |
| structure | A structure
|
Example: "Prior",struct("ClassNames",["b","g"],"ClassProbs",1:2)
Data Types: single | double | char | string | struct
Response variable name, specified as a character vector or string scalar.
If you supply
Y, then you can useResponseNameto specify a name for the response variable.If you supply
ResponseVarNameorformula, then you cannot useResponseName.
Example: ResponseName="response"
Data Types: char | string
Observation weights, specified as a positive numeric vector or the name of a
variable in Tbl. The software weights each observation in
X or Tbl with the corresponding value in
Weights. The length of Weights must equal
the number of rows in X or Tbl.
If you specify the input data as a table Tbl, then
Weights can be the name of a variable in
Tbl that contains a numeric vector. In this case, you must
specify Weights as a character vector or string scalar. For
example, if the weights vector W is stored as
Tbl.W, then specify it as "W". Otherwise, the
software treats all columns of Tbl, including
W, as predictors or the response variable when training the
model.
By default, Weights is ones(n,1), where
n is the number of observations in X or
Tbl.
The software normalizes Weights to sum to the value of the
prior probability in the respective class. Inf weights are not supported.
Data Types: single | double | char | string
Output Arguments
Trained classification model, returned as one of the classification model objects in the following table or a cell array of model objects.
| Learner Name | Returned Model Object |
|---|---|
"discr" | CompactClassificationDiscriminant |
"ensemble" |
|
"kernel" |
|
"knn" | ClassificationKNN |
"linear" |
|
"nb" | CompactClassificationNaiveBayes |
"net" | CompactClassificationNeuralNetwork |
"svm" |
|
"tree" | CompactClassificationTree |
If you specify OptimizeHyperparameters and
set the ConstraintType and ConstraintBounds options of
HyperparameterOptimizationOptions, then Mdl is an
N-by-1 cell array of model objects, where N is equal
to the number of rows in ConstraintBounds. If none of the optimization
problems yields a feasible model, then each cell array value is [].
Optimization results, returned as a SupervisedLearningBayesianOptimization object, an AggregateBayesianOptimization object, or a table. If you set the
ConstraintType and ConstraintBounds options
of HyperparameterOptimizationOptions, then
OptimizationResults is an
AggregateBayesianOptimization object. Otherwise,
OptimizationResults is a
SupervisedLearningBayesianOptimization object if
Optimizer is "bayesopt", or a table if
Optimizer is "asha". For more information, see
Bayesian Optimization and ASHA Optimization.
More About
When you set the Verbose field of the
HyperparameterOptimizationOptions name-value argument to
1 or 2, the fitcauto function
provides an iterative display of the optimization results.
The following table describes the columns in the display and their entries.
| Column Name | Description |
|---|---|
Iter | Iteration number — You can set a limit to the number of iterations by using
the MaxObjectiveEvaluations field of the
HyperparameterOptimizationOptions name-value
argument. |
Active workers | Number of active parallel workers — This column appears only when you run the
optimization in parallel by setting the UseParallel field of
the HyperparameterOptimizationOptions name-value argument to
true. |
Eval result | One of the following evaluation results:
|
Validation loss | Validation loss computed for the learner and hyperparameter values at
this iteration — In particular, You can
change the validation scheme by using the |
Time for training & validation (sec) | Time taken to train and compute the validation loss for the model with the learner and hyperparameter values at this iteration (in seconds) — When you use Bayesian optimization, this value excludes the time required to update the objective function model maintained by the Bayesian optimization process. For more details, see Bayesian Optimization. |
Observed min validation loss | Observed minimum validation loss computed so far — This value
corresponds to the smallest By default,
|
Estimated min validation loss | Estimated minimum validation loss — When you use Bayesian optimization,
By default, Note This column appears only when you use Bayesian optimization, that is, when
the |
Training set size | Number of observations used in each training set at this iteration —
Use the Note This column appears only when you use ASHA optimization, that is, when the
|
Constraint1 violation | Amount by which the constraint bounds are violated. A nonpositive value
indicates that the value of the constraint is within the constraint bounds. This
column is displayed if you set the ConstraintType and
ConstraintBounds options of HyperparameterOptimizationOptions. |
Learner | Model type evaluated at this iteration — Specify the learners used in the
optimization by using the Learners name-value
argument. |
Hyperparameter: Value | Hyperparameter values at this iteration — Specify the hyperparameters used in
the optimization by using the OptimizeHyperparameters
name-value argument. |
The display also includes these model descriptions:
Best observed learner— This model, with the listed learner type and hyperparameter values, yields the final observed minimum validation loss. When you use ASHA optimization,fitcautoretrains the model on the entire training data set and returns it as theMdloutput.Best estimated learner— This model, with the listed learner type and hyperparameter values, yields the final estimated minimum validation loss when you use Bayesian optimization. In this case,fitcautoretrains the model on the entire training data set and returns it as theMdloutput.Note
The
Best estimated learnermodel appears only when you use Bayesian optimization, that is, when theOptimizerfield of theHyperparameterOptimizationOptionsname-value argument is set to"bayesopt".
Tips
Depending on the size of your data set, the number of learners you specify, and the optimization method you choose,
fitcautocan take some time to run.If you have a Parallel Computing Toolbox license, you can speed up computations by running the optimization in parallel. To do so, specify
"HyperparameterOptimizationOptions",struct("UseParallel",true). You can include additional fields in the structure to control other aspects of the optimization. SeeHyperparameterOptimizationOptions.If
fitcautowith Bayesian optimization takes a long time to run because of the number of observations in your training set (for example, over 10,000), consider usingfitcautowith ASHA optimization instead. ASHA optimization often finds good solutions faster than Bayesian optimization for data sets with many observations. To use ASHA optimization, specify"HyperparameterOptimizationOptions",struct("Optimizer","asha"). You can include additional fields in the structure to control other aspects of the optimization. In particular, if you have a time constraint, specify theMaxTimefield of theHyperparameterOptimizationOptionsstructure to limit the number of secondsfitcautoruns.
Classifiers use different criteria to predict labels. For example, some classifiers predict labels using the maximum score while others use the minimum expected cost.
fitcautodoes not consider how models predict labels when selecting the best model. To see how the returned classifierMdlclassifies new observations, consult the correspondingpredictfunction.
Algorithms
When you specify "Learners","auto", the fitcauto
function analyzes the predictor and response data in order to choose appropriate learners.
The function considers whether the data set has any of these characteristics:
Categorical predictors
Missing values for more than 5% of the data
Imbalanced data, where the ratio of the number of observations in the largest class to the number of observations in the smallest class is greater than 5
More than 100 observations in the smallest class
Wide data, where the number of predictors is greater than or equal to the number of observations
High-dimensional data, where the number of predictors is greater than 100
Large data, where the number of observations is greater than 50,000
Binary response variable
Ordinal response variable
The selected learners are always a subset of those listed in the
Learners table. However, the associated models tried during the
optimization process can have different default values for hyperparameters not being
optimized, as well as different search ranges for hyperparameters being optimized.
The goal of Bayesian optimization, and optimization in general, is to find a point that
minimizes an objective function. In the context of fitcauto, a point is
a learner type together with a set of hyperparameter values for the learner (see Learners and
OptimizeHyperparameters), and the objective function is the cross-validation
classification error, by default. The Bayesian optimization implemented in
fitcauto internally maintains a multi-TreeBagger model of the objective function. That is, the objective function
model splits along the learner type and, for a given learner, the model is a
TreeBagger ensemble for regression. (This underlying model differs from
the Gaussian process model employed by other Statistics and Machine Learning Toolbox™ functions that use Bayesian optimization.) Bayesian optimization trains the
underlying model by using objective function evaluations, and determines the next point to
evaluate by using an acquisition function ("expected-improvement"). For
more information, see Expected Improvement. The acquisition function balances between sampling at
points with low modeled objective function values and exploring areas that are not well
modeled yet. At the end of the optimization, fitcauto chooses the point
with the minimum objective function model value, among the points evaluated during the
optimization. For more information, see the
"Criterion","min-visited-mean" name-value argument of bestPoint.
The asynchronous successive halving algorithm (ASHA) in fitcauto
randomly chooses several models with different hyperparameter values (see Learners and
OptimizeHyperparameters) and trains them on a small subset of the training
data. If the performance of a particular model is promising, the model is promoted and
trained on a larger amount of the training data. This process repeats, and successful models
are trained on progressively larger amounts of data. By default, at the end of the
optimization, fitcauto chooses the model that has the lowest
cross-validation classification error.
At each iteration, ASHA either chooses a previously trained model and promotes it (that is, retrains the model using more training data), or selects a new model (learner type and hyperparameter values) using random search. ASHA promotes models as follows:
The algorithm searches for the group of models with the largest training set size for which this condition does not hold:
floor(g/4)of the models have been promoted, wheregis the number of models in the group.Among the group of models, ASHA chooses the model with the lowest cross-validation classification error and retrains that model with
4*(Training Set Size)observations.If no such group of models exists, then ASHA selects a new model instead of promoting an old one, and trains the new model using the smallest training set size.
When a model is trained on a subset of the training data, ASHA computes the cross-validation classification error as follows:
For each training fold, the algorithm selects a random sample of the observations (of size
Training set size) using stratified sampling, and then trains a model on that subset of data.The algorithm then tests the fitted model on the test fold (that is, the observations not in the training fold) and computes the classification error.
Finally, the algorithm averages the results across all folds.
For more information on ASHA, see [1].
When you use ASHA optimization, the default number of iterations depends on the number
of observations in the data, the number of learner types, the use of parallel processing,
and the type of cross-validation. The algorithm selects the number of iterations such that,
for L learner types (see Learners),
fitcauto trains L models on the largest training
set size.
This table describes the default number of iterations based on the given specifications when you use 5-fold cross-validation. Note that n represents the number of observations and L represents the number of learner types.
Number of Observations n | Default Number of Iterations (run in serial) | Default Number of Iterations (run in parallel) |
|---|---|---|
| n < 500 | 30*L — n is too small to implement ASHA
optimization, and fitcauto implements random search to find and
assess models instead. | 30*L — n is too small to implement ASHA
optimization, and fitcauto implements random search to find and
assess models instead. |
| 500 ≤ n < 2000 | 5*L | 5*(L + 1) |
| 2000 ≤ n < 8000 | 21*L | 21*(L + 1) |
| 8000 ≤ n < 32,000 | 85*L | 85*(L + 1) |
| 32,000 ≤ n | 341*L | 341*(L + 1) |
Alternative Functionality
If you are unsure which models work best for your data set, you can alternatively use the Classification Learner app. Using the app, you can perform hyperparameter tuning for different models, and choose the optimized model that performs best. Although you must select a specific model before you can tune the model hyperparameters, Classification Learner provides greater flexibility for selecting optimizable hyperparameters and setting hyperparameter values. However, you cannot optimize in parallel, specify observation weights, specify prior probabilities, or use ASHA optimization in the app. For more information, see Hyperparameter Optimization in Classification Learner App.
If you know which models might suit your data, you can alternatively use the corresponding model fit functions and specify the
OptimizeHyperparametersname-value argument to tune hyperparameters. You can compare the results across the models to select the best classifier. For an example of this process, see Moving Towards Automating Model Selection Using Bayesian Optimization.
References
[1] Li, Liam, Kevin Jamieson, Afshin Rostamizadeh, Ekaterina Gonina, Moritz Hardt, Benjamin Recht, and Ameet Talwalkar. “A System for Massively Parallel Hyperparameter Tuning.” ArXiv:1810.05934v5 [Cs], March 16, 2020. https://arxiv.org/abs/1810.05934v5.
Extended Capabilities
To perform parallel hyperparameter optimization, use the
UseParallel=true option in the
HyperparameterOptimizationOptions name-value pair argument in the call
to the fitcauto function.
For more information on parallel hyperparameter optimization, see Parallel Bayesian Optimization.
For general information about parallel computing, see Run MATLAB Functions with Automatic Parallel Support (Parallel Computing Toolbox).
Version History
Introduced in R2020aYou can choose when to select models and their hyperparameters with respect to
misclassification cost. In the HyperparameterOptimizationOptions
structure or object, set the LossFun value to
"auto-cost" if you want to optimize using the appropriate
cost-sensitive loss for each model. Alternatively, set the LossFun value
to "classiferror" to optimize using the misclassified rate in decimal. In
previous releases, the function always uses the unweighted mean misclassification cost when
you specify the Cost name-value argument.
If you perform Bayesian optimization, you can return the optimization results
(OptimizationResults), which are stored in a SupervisedLearningBayesianOptimization object. In previous releases, the
optimization results are stored in a BayesianOptimization object.
fitcauto defaults to serial hyperparameter optimization when
HyperparameterOptimizationOptions includes
UseParallel=true and the software cannot open a parallel pool.
In previous releases, the software issues an error under these circumstances.
Starting in R2024b, fitcauto includes ensembles with
"AdaBoostM1", "GentleBoost", and
"LogitBoost" ensemble methods as part of the model selection and
hyperparameter tuning process for multiclass classification. For such models,
fitcauto additionally tunes the coding design hyperparameter
(Coding). If the function selects
one of these models as the best estimated learner, the returned model is a CompactClassificationECOC model object.
The selection of ensemble models differs from previous releases only when the response
variable contains three or more classes, and the Learners value is
"auto", "all", "all-nonlinear",
or a string or cell array containing the "ensemble" learner name.
Starting in R2023b, when the Learners value includes kernel
("kernel"), k-nearest neighbor
("knn"), naive Bayes ("nb"), or support vector
machine ("svm") classifiers, the fitcauto
function optimizes the Standardize hyperparameter of the models by
default. That is, if the OptimizeHyperparameters value is
"auto", then Standardize is an optimizable
hyperparameter of kernel, KNN, naive Bayes, and SVM models.
Starting in R2023b, fitcauto optimizes the kernel smoothing window width of naive Bayes models by using the default search range [1e-3,1e3]. That is, when you specify to optimize the naive Bayes hyperparameter Width by using the OptimizeHyperparameters name-value argument, the function searches among positive values log-scaled in the range [1e-3,1e3].
In previous releases, the default search range for the Width
hyperparameter was
[MinPredictorDiff/4,max(MaxPredictorRange,MinPredictorDiff)], where
MinPredictorDiff and MaxPredictorRange were
determined as follows:
diffs = diff(sort(X));
MinPredictorDiff = min(diffs(diffs ~= 0),[],"omitnan");
MaxPredictorRange = max(max(X) - min(X));Starting in R2023a, fitcauto supports misclassification costs and
prior probabilities for neural network classifiers. That is, you can specify the
Cost and Prior name-value arguments when the
Learners name-value argument includes "net"
models.
In previous releases, when you specified nondefault misclassification costs or prior
probabilities, fitcauto omitted neural network models from the model
selection process.
Starting in R2022a, the list of available learners includes neural network models. When you
specify "all" or "all-nonlinear" for the
Learners name-value argument, fitcauto
includes neural network models as part of the model selection and hyperparameter tuning
process. The function also considers neural network models when you specify
Learners as "auto", depending on the
characteristics of your data set.
To omit neural network models from the model selection process, you can explicitly specify the
models you want to include. For example, to use tree and ensemble models only, specify
"Learners",["tree","ensemble"].
Starting in R2022a, if you specify Learners as
"auto" and the data has more predictors than observations after the
expansion of the categorical predictors (see Automatic Creation of Dummy Variables), then
fitcauto includes linear learners ("linear")
along with other models during the hyperparameter optimization. In previous releases, linear
learners were not considered.
Starting in R2022a, when you specify to try a linear learner
("linear") for multiclass classification,
fitcauto uses either a Limited-memory BFGS (LBFGS) solver or a
Sparse Reconstruction by Separable Approximation (SpaRSA) solver, depending on the
regularization type selected during that iteration of the optimization process.
When
Regularizationis'ridge', the function sets theSolvervalue to'lbfgs'by default.When
Regularizationis'lasso', the function sets theSolvervalue to'sparsa'by default.
In previous releases, the default solver selection during the optimization process
depended on various factors, including the regularization type, learner type, and number of
predictors. For more information, see Solver.
Starting in R2021a, when you specify to try a linear learner
("linear") for binary classification, fitcauto
uses either a Limited-memory BFGS (LBFGS) solver or a Sparse Reconstruction by Separable
Approximation (SpaRSA) solver, depending on the regularization type selected during that
iteration of the optimization process.
When
Regularizationis'ridge', the function sets theSolvervalue to'lbfgs'by default.When
Regularizationis'lasso', the function sets theSolvervalue to'sparsa'by default.
In previous releases, the default solver selection during the optimization process
depended on various factors, including the regularization type, learner type, and number of
predictors. For more information, see Solver.
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)