Image Captioning Using Attention
This example shows how to train a deep learning model for image captioning using attention.
Most pretrained deep learning networks are configured for single-label classification. For example, given an image of a typical office desk, the network might predict the single class "keyboard" or "mouse". In contrast, an image captioning model combines convolutional and recurrent operations to produce a textual description of what is in the image, rather than a single label.
This model trained in this example uses an encoder-decoder architecture. The encoder is a pretrained Inception-v3 network used as a feature extractor. The decoder is a recurrent neural network (RNN) that takes the extracted features as input and generates a caption. The decoder incorporates an attention mechanism that allows the decoder to focus on parts of the encoded input while generating the caption.

The encoder model is a pretrained Inception-v3 model that extracts features from the "mixed10" layer, followed by fully connected and ReLU operations.

The decoder model consists of a word embedding, an attention mechanism, a gated recurrent unit (GRU), and two fully connected operations.

Load Pretrained Network
Load a pretrained Incetion-v3 network. This step requires the Deep Learning Toolbox™ Model for Inception-v3 Network support package. If you do not have the required support package installed, then the software provides a download link.
net = imagePretrainedNetwork("inceptionv3");
inputSizeNet = net.Layers(1).InputSize;Remove the last three layers, leaving the "mixed10" layer as the last layer.
net = removeLayers(net, ["avg_pool" "predictions" "predictions_softmax"]);
View the input layer of the network. The Inception-v3 network uses symmetric-rescale normalization with a minimum value of 0 and a maximum value of 255.
net.Layers(1)
ans =
ImageInputLayer with properties:
Name: 'input_1'
InputSize: [299 299 3]
SplitComplexInputs: 0
Hyperparameters
DataAugmentation: 'none'
Normalization: 'rescale-symmetric'
NormalizationDimension: 'auto'
Max: 255
Min: 0
Custom training does not support this normalization, so you must disable normalization in the network and perform the normalization in the custom training loop instead. Save the minimum and maximum values as doubles in variables named inputMin and inputMax, respectively, and replace the input layer with an image input layer without normalization.
inputMin = double(net.Layers(1).Min); inputMax = double(net.Layers(1).Max); layer = imageInputLayer(inputSizeNet,Normalization="none",Name="input"); net = replaceLayer(net,"input_1",layer);
Initialize the network.
net = initialize(net);
Determine the output size of the network. Use the analyzeNetwork function to see the activation sizes of the last layer.
analyzeNetwork(net)
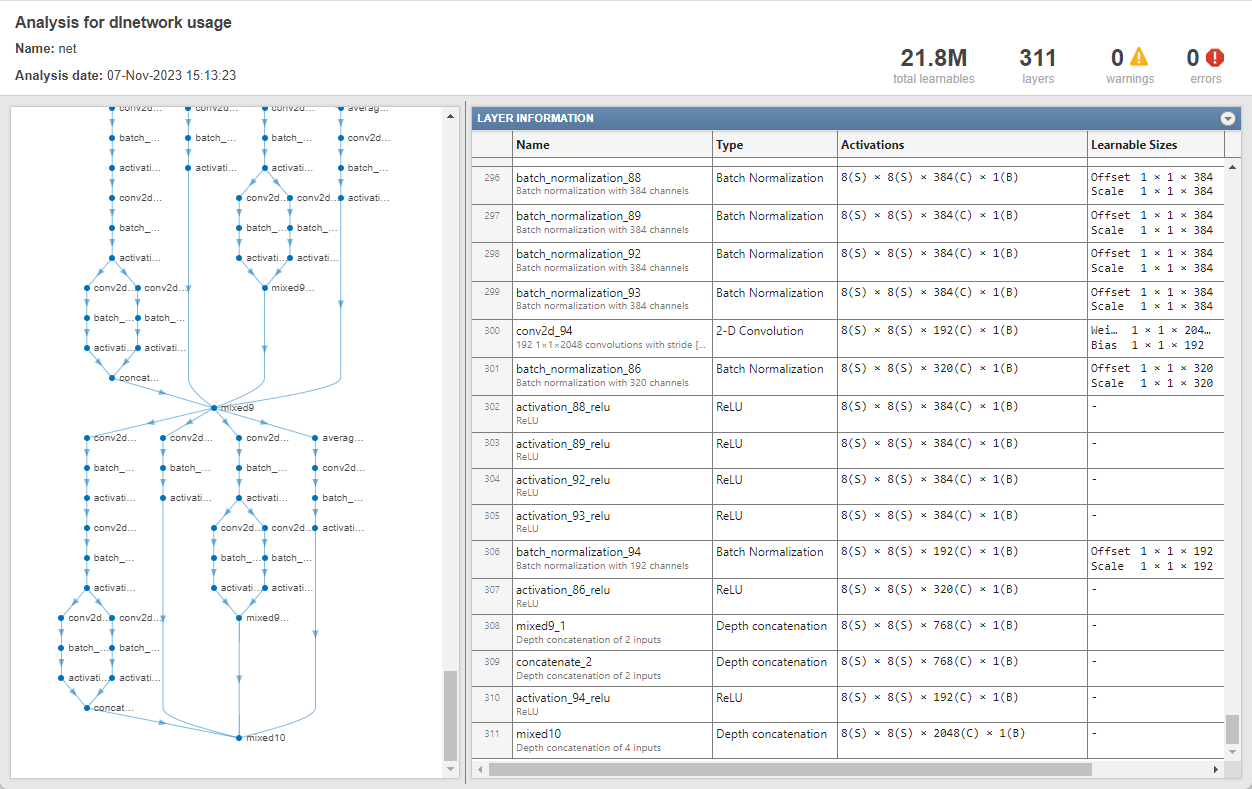
Create a variable named outputSizeNet containing the network output size.
outputSizeNet = [8 8 2048];
Import COCO Data Set
Download images and annotations from the data sets "2014 Train images" and "2014 Train/val annotations," respectively, from https://cocodataset.org/#download. Extract the images and annotations into a folder named "coco". The COCO 2014 data set was collected by Coco Consortium.
Extract the captions from the file "captions_train2014.json" using the jsondecode function.
dataFolder = fullfile(tempdir,"coco"); filename = fullfile(dataFolder,"annotations_trainval2014","annotations","captions_train2014.json"); str = fileread(filename); data = jsondecode(str)
data = struct with fields:
info: [1×1 struct]
images: [82783×1 struct]
licenses: [8×1 struct]
annotations: [414113×1 struct]
The annotations field of the struct contains the data required for image captioning.
data.annotations
ans=414113×1 struct array with fields:
318556 48 'A very clean and well decorated empty bathroom'
116100 67 'A panoramic view of a kitchen and all of its appliances.'
318556 126 'A blue and white bathroom with butterfly themed wall tiles.'
116100 148 'A panoramic photo of a kitchen and dining room'
379340 173 'A graffiti-ed stop sign across the street from a red car '
379340 188 'A vandalized stop sign and a red beetle on the road'
318556 219 'A bathroom with a border of butterflies and blue paint on the walls above it.'
318556 255 'An angled view of a beautifully decorated bathroom.'
134754 272 'The two people are walking down the beach.'
538480 288 'A sink and a toilet inside a small bathroom.'
476220 314 'An empty kitchen with white and black appliances.'
299675 328 'A white square kitchen with tile floor that needs repairs '
32275 352 'The vanity contains two sinks with a towel for each.'
302443 411 'Several metal balls sit in the sand near a group of people.'
The data set contains multiple captions for each image. To ensure the same images do not appear in both training and validation sets, identify the unique images in the data set using the unique function by using the IDs in the image_id field of the annotations field of the data, then view the number of unique images.
numObservationsAll = numel(data.annotations)
numObservationsAll = 414113
imageIDs = [data.annotations.image_id]; imageIDsUnique = unique(imageIDs); numUniqueImages = numel(imageIDsUnique)
numUniqueImages = 82783
Each image has at least five captions. Create a struct annotationsAll with these fields:
ImageID— Image IDFilename— File name of the imageCaptions— String array of raw captionsCaptionIDs— Vector of indices of the corresponding captions indata.annotations
To make merging easier, sort the annotations by the image IDs.
[~,idx] = sort([data.annotations.image_id]); data.annotations = data.annotations(idx);
Loop over the annotations and merge multiple annotations when necessary.
i = 0; j = 0; imageIDPrev = 0; while i < numel(data.annotations) i = i + 1; imageID = data.annotations(i).image_id; caption = string(data.annotations(i).caption); if imageID ~= imageIDPrev % Create new entry j = j + 1; annotationsAll(j).ImageID = imageID; annotationsAll(j).Filename = fullfile(dataFolder,"train2014","COCO_train2014_" + pad(string(imageID),12,"left","0") + ".jpg"); annotationsAll(j).Captions = caption; annotationsAll(j).CaptionIDs = i; else % Append captions annotationsAll(j).Captions = [annotationsAll(j).Captions; caption]; annotationsAll(j).CaptionIDs = [annotationsAll(j).CaptionIDs; i]; end imageIDPrev = imageID; end
Partition the data into training and validation sets. Hold out 5% of the observations for testing.
cvp = cvpartition(numel(annotationsAll),HoldOut=0.05); idxTrain = training(cvp); idxTest = test(cvp); annotationsTrain = annotationsAll(idxTrain); annotationsTest = annotationsAll(idxTest);
The struct contains three fields:
id— Unique identifier for the captioncaption— Image caption, specified as a character vectorimage_id— Unique identifier of the image corresponding to the caption
To view the image and the corresponding caption, locate the image file with file name "train2014\COCO_train2014_XXXXXXXXXXXX.jpg", where "XXXXXXXXXXXX" corresponds to the image ID left-padded with zeros to have length 12.
imageID = annotationsTrain(1).ImageID; captions = annotationsTrain(1).Captions; filename = annotationsTrain(1).Filename;
To view the image, use the imread and imshow functions.
img = imread(filename); figure imshow(img) title(captions)
Prepare Data for Training
Prepare the captions for training and testing. Extract the text from the Captions field of the struct containing both the training and test data (annotationsAll), erase the punctuation, and convert the text to lowercase.
captionsAll = cat(1,annotationsAll.Captions); captionsAll = erasePunctuation(captionsAll); captionsAll = lower(captionsAll);
In order to generate captions, the RNN decoder requires special start and stop tokens to indicate when to start and stop generating text, respectively. Add the custom tokens "<start>" and "<stop>" to the beginnings and ends of the captions, respectively.
captionsAll = "<start>" + captionsAll + "<stop>";
Tokenize the captions using the tokenizedDocument function and specify the start and stop tokens using the CustomTokens option.
documentsAll = tokenizedDocument(captionsAll,CustomTokens=["<start>" "<stop>"]);
Create a wordEncoding object that maps words to numeric indices and back. Reduce the memory requirements by specifying a vocabulary size of 5000 corresponding to the most frequently observed words in the training data. To avoid bias, use only the documents corresponding to the training set.
enc = wordEncoding(documentsAll(idxTrain),MaxNumWords=5000,Order="frequency");Create an augmented image datastore containing the images corresponding to the captions. Set the output size to match the input size of the convolutional network. To keep the images synchronized with the captions, specify a table of file names for the datastore by reconstructing the file names using the image ID. To return grayscale images as 3-channel RGB images, set the ColorPreprocessing option to "gray2rgb".
tblFilenames = table(cat(1,annotationsTrain.Filename));
augimdsTrain = augmentedImageDatastore(inputSizeNet,tblFilenames,ColorPreprocessing="gray2rgb")augimdsTrain =
augmentedImageDatastore with properties:
NumObservations: 78644
MiniBatchSize: 1
DataAugmentation: 'none'
ColorPreprocessing: 'gray2rgb'
OutputSize: [299 299]
OutputSizeMode: 'resize'
DispatchInBackground: 0
Initialize Model Parameters
Initialize the model parameters. Specify 512 hidden units with a word embedding dimension of 256.
embeddingDimension = 256; numHiddenUnits = 512;
Initialize a struct containing the parameters for the encoder model.
Initialize the weights of the fully connected operations using the Glorot initializer, specified by the
initializeGlorotfunction, listed at the end of the example. Specify the output size to match the embedding dimension of the decoder (256) and an input size to match the number of output channels of the pretrained network. The'mixed10'layer of the Inception-v3 network outputs data with 2048 channels.
numFeatures = outputSizeNet(1) * outputSizeNet(2); inputSizeEncoder = outputSizeNet(3); parametersEncoder = struct; % Fully connect parametersEncoder.fc.Weights = dlarray(initializeGlorot(embeddingDimension,inputSizeEncoder)); parametersEncoder.fc.Bias = dlarray(zeros([embeddingDimension 1],"single"));
Initialize a struct containing parameters for the decoder model.
Initialize the word embedding weights with the size given by the embedding dimension and the vocabulary size plus one, where the extra entry corresponds to the padding value.
Initialize the weights and biases for the Bahdanau attention mechanism with sizes corresponding to the number of hidden units of the GRU operation.
Initialize the weights and bias of the GRU operation.
Initialize the weights and biases of two fully connected operations.
For the model decoder parameters, initialize each of the weighs and biases with the Glorot initializer and zeros, respectively.
inputSizeDecoder = enc.NumWords + 1; parametersDecoder = struct; % Word embedding parametersDecoder.emb.Weights = dlarray(initializeGlorot(embeddingDimension,inputSizeDecoder)); % Attention parametersDecoder.attention.Weights1 = dlarray(initializeGlorot(numHiddenUnits,embeddingDimension)); parametersDecoder.attention.Bias1 = dlarray(zeros([numHiddenUnits 1],"single")); parametersDecoder.attention.Weights2 = dlarray(initializeGlorot(numHiddenUnits,numHiddenUnits)); parametersDecoder.attention.Bias2 = dlarray(zeros([numHiddenUnits 1],"single")); parametersDecoder.attention.WeightsV = dlarray(initializeGlorot(1,numHiddenUnits)); parametersDecoder.attention.BiasV = dlarray(zeros(1,1,"single")); % GRU parametersDecoder.gru.InputWeights = dlarray(initializeGlorot(3*numHiddenUnits,2*embeddingDimension)); parametersDecoder.gru.RecurrentWeights = dlarray(initializeGlorot(3*numHiddenUnits,numHiddenUnits)); parametersDecoder.gru.Bias = dlarray(zeros(3*numHiddenUnits,1,"single")); % Fully connect parametersDecoder.fc1.Weights = dlarray(initializeGlorot(numHiddenUnits,numHiddenUnits)); parametersDecoder.fc1.Bias = dlarray(zeros([numHiddenUnits 1],"single")); % Fully connect parametersDecoder.fc2.Weights = dlarray(initializeGlorot(enc.NumWords+1,numHiddenUnits)); parametersDecoder.fc2.Bias = dlarray(zeros([enc.NumWords+1 1],"single"));
Define Model Functions
Create the functions modelEncoder and modelDecoder, listed at the end of the example, which compute the outputs of the encoder and decoder models, respectively.
The modelEncoder function, listed in the Encoder Model Function section of the example, takes as input an array of activations X from the output of the pretrained network and passes it through a fully connected operation and a ReLU operation. Because the pretrained network does not need to be traced for automatic differentiation, extracting the features outside the encoder model function is more computationally efficient.
The modelDecoder function, listed in the Decoder Model Function section of the example, takes as input a single input time-step corresponding to an input word, the decoder model parameters, the features from the encoder, and the network state, and returns the predictions for the next time step, the updated network state, and the attention weights.
Specify Training Options
Specify the options for training. Train for 30 epochs with a mini-batch size of 128 and display the training progress in a plot.
miniBatchSize = 128;
numEpochs = 30;
plots = "training-progress";Train on a GPU if one is available. Using a GPU requires Parallel Computing Toolbox™ and a supported GPU device. For information on supported devices, see GPU Computing Requirements (Parallel Computing Toolbox).
executionEnvironment = "auto";Check whether a GPU is available for training.
if canUseGPU gpu = gpuDevice; disp(gpu.Name + " GPU detected and available for training.") end
NVIDIA RTX A5000 GPU detected and available for training.
Train Network
Train the network using a custom training loop.
At the beginning of each epoch, shuffle the input data. To keep the images in the augmented image datastore and the captions synchronized, create an array of shuffled indices that indexes into both data sets.
For each mini-batch:
Rescale the images to the size that the pretrained network expects.
For each image, select a random caption.
Convert the captions to sequences of word indices. Specify right-padding of the sequences with the padding value corresponding to the index of the padding token.
Convert the data to
dlarrayobjects. For the images, specify dimension labels"SSCB"(spatial, spatial, channel, batch).For GPU training, convert the data to
gpuArrayobjects.Extract the image features using the pretrained network and reshape them to the size the encoder expects.
Evaluate the model loss and gradients using the
dlfevalandmodelLossfunctions.Update the encoder and decoder model parameters using the
adamupdatefunction.Display the training progress in a plot.
Initialize the parameters for the Adam optimizer.
trailingAvgEncoder = []; trailingAvgSqEncoder = []; trailingAvgDecoder = []; trailingAvgSqDecoder = [];
Initialize the TrainingProgressMonitor object. Because the timer starts when you create the monitor object, make sure that you create the object close to the training loop.
if plots == "training-progress" monitor = trainingProgressMonitor( ... Metrics="Loss", ... Info="Epoch", ... XLabel="Iteration"); end
Train the model.
iteration = 0; numObservationsTrain = numel(annotationsTrain); numIterationsPerEpoch = floor(numObservationsTrain / miniBatchSize); numIterations = numIterationsPerEpoch*numEpochs; % Loop over epochs. for epoch = 1:numEpochs % Shuffle data. idxShuffle = randperm(numObservationsTrain); % Loop over mini-batches. for i = 1:numIterationsPerEpoch iteration = iteration + 1; % Determine mini-batch indices. idx = (i-1)*miniBatchSize+1:i*miniBatchSize; idxMiniBatch = idxShuffle(idx); % Read mini-batch of data. tbl = readByIndex(augimdsTrain,idxMiniBatch); X = cat(4,tbl.input{:}); annotations = annotationsTrain(idxMiniBatch); % For each image, select random caption. idx = cellfun(@(captionIDs) randsample(captionIDs,1),{annotations.CaptionIDs}); documents = documentsAll(idx); % Create batch of data. [X,T] = createBatch(X,documents,net,inputMin,inputMax,enc,executionEnvironment); % Evaluate the model loss and gradients using dlfeval and the % modelLoss function. [loss,gradientsEncoder,gradientsDecoder] = dlfeval(@modelLoss,parametersEncoder, ... parametersDecoder,X,T); % Update encoder using adamupdate. [parametersEncoder,trailingAvgEncoder,trailingAvgSqEncoder] = adamupdate(parametersEncoder, ... gradientsEncoder,trailingAvgEncoder,trailingAvgSqEncoder,iteration); % Update decoder using adamupdate. [parametersDecoder,trailingAvgDecoder,trailingAvgSqDecoder] = adamupdate(parametersDecoder, ... gradientsDecoder,trailingAvgDecoder,trailingAvgSqDecoder,iteration); % Display the training progress. if plots == "training-progress" recordMetrics(monitor,iteration,Loss=loss); updateInfo(monitor,Epoch=epoch); monitor.Progress = 100 * iteration/numIterations; end end end

Predict New Captions
The caption generation process is different from the process for training. During training, at each time step, the decoder uses the true value of the previous time step as input. This is known as "teacher forcing". When making predictions on new data, the decoder uses the previous predicted values instead of the true values.
Predicting the most likely word for each step in the sequence can lead to suboptimal results. For example, if the decoder predicts the first word of a caption is "a" when given an image of an elephant, then the probability of predicting "elephant" for the next word becomes much more unlikely because of the extremely low probability of the phrase "a elephant" appearing in English text.
To address this issue, you can use the beam search algorithm: instead of taking the most likely prediction for each step in the sequence, take the top k predictions (the beam index) and for each following step, keep the top k predicted sequences so far according to the overall score.
Generate a caption of a new image by extracting the image features, inputting them into the encoder, and then using the beamSearch function, listed in the Beam Search Function section of the example.
img = imread("dog_sitting.jpg");
X = extractImageFeatures(net,img,inputMin,inputMax,executionEnvironment);
beamIndex = 3;
maxNumWords = 20;
[words,attentionScores] = beamSearch(X,beamIndex,parametersEncoder,parametersDecoder,enc,maxNumWords);
caption = join(words)caption = "a small white dog standing on a lush green grass covered field"
Display the image with the caption.
figure imshow(img) title(caption)

Predict Captions for Data Set
To predict captions for a collection of images, loop over mini-batches of data in the datastore and extract the features from the images using the extractImageFeatures function. Then, loop over the images in the mini-batch and generate captions using the beamSearch function.
Create an augmented image datastore and set the output size to match the input size of the convolutional network. To output grayscale images as 3-channel RGB images, set the ColorPreprocessing option to "gray2rgb".
tblFilenamesTest = table(cat(1,annotationsTest.Filename));
augimdsTest = augmentedImageDatastore(inputSizeNet,tblFilenamesTest,ColorPreprocessing="gray2rgb")augimdsTest =
augmentedImageDatastore with properties:
NumObservations: 4139
MiniBatchSize: 1
DataAugmentation: 'none'
ColorPreprocessing: 'gray2rgb'
OutputSize: [299 299]
OutputSizeMode: 'resize'
DispatchInBackground: 0
Generate captions for the test data. Predicting captions on a large data set can take some time. If you have Parallel Computing Toolbox™, then you can make predictions in parallel by generating captions inside a parfor loop. If you do not have Parallel Computing Toolbox. then the parfor loop runs in serial.
beamIndex = 2; maxNumWords = 20; numObservationsTest = numel(annotationsTest); numIterationsTest = ceil(numObservationsTest/miniBatchSize); captionsTestPred = strings(1,numObservationsTest); documentsTestPred = tokenizedDocument(strings(1,numObservationsTest)); for i = 1:numIterationsTest % Mini-batch indices. idxStart = (i-1)*miniBatchSize+1; idxEnd = min(i*miniBatchSize,numObservationsTest); idx = idxStart:idxEnd; sz = numel(idx); % Read images. tbl = readByIndex(augimdsTest,idx); % Extract image features. X = cat(4,tbl.input{:}); X = extractImageFeatures(net,X,inputMin,inputMax,executionEnvironment); % Generate captions. captionsPredMiniBatch = strings(1,sz); documentsPredMiniBatch = tokenizedDocument(strings(1,sz)); parfor j = 1:sz words = beamSearch(X(:,:,j),beamIndex,parametersEncoder,parametersDecoder,enc,maxNumWords); captionsPredMiniBatch(j) = join(words); documentsPredMiniBatch(j) = tokenizedDocument(words,TokenizeMethod="none"); end captionsTestPred(idx) = captionsPredMiniBatch; documentsTestPred(idx) = documentsPredMiniBatch; end
To view a test image with the corresponding caption, use the imshow function and set the title to the predicted caption.
idx = 1;
tbl = readByIndex(augimdsTest,idx);
img = tbl.input{1};
figure
imshow(img)
title(captionsTestPred(idx))
Evaluate Model Accuracy
To evaluate the accuracy of the captions using the BLEU score, calculate the BLEU score for each caption (the candidate) against the corresponding captions in the test set (the references) using the bleuEvaluationScore function. Using the bleuEvaluationScore function, you can compare a single candidate document to multiple reference documents.
The bleuEvaluationScore function, by default, scores similarity using n-grams of length one through four. As the captions are short, this behavior can lead to uninformative results as most scores are close to zero. Set the n-gram length to one through two by setting the NgramWeights option to a two-element vector with equal weights.
ngramWeights = [0.5 0.5]; for i = 1:numObservationsTest annotation = annotationsTest(i); captionIDs = annotation.CaptionIDs; candidate = documentsTestPred(i); references = documentsAll(captionIDs); score = bleuEvaluationScore(candidate,references,NgramWeights=ngramWeights); scores(i) = score; end
View the mean BLEU score.
scoreMean = mean(scores)
scoreMean = 0.3875
Visualize the scores in a histogram.
figure histogram(scores) xlabel("BLEU Score") ylabel("Frequency")
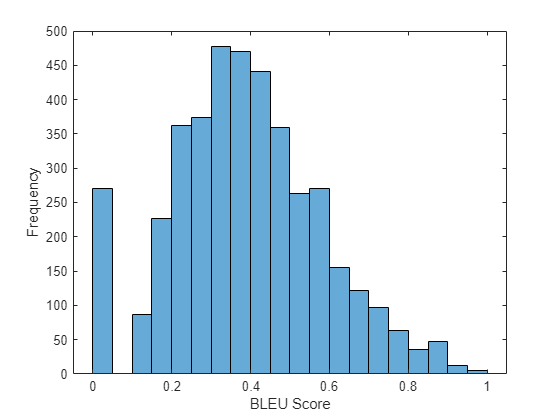
Attention Function
The attention function calculates the context vector and the attention weights using Bahdanau attention.
function [contextVector, attentionWeights] = attention(hidden,features,weights1, ... bias1,weights2,bias2,weightsV,biasV) % Model dimensions. [embeddingDimension,numFeatures,miniBatchSize] = size(features); numHiddenUnits = size(weights1,1); % Fully connect. Y1 = reshape(features,embeddingDimension, numFeatures*miniBatchSize); Y1 = fullyconnect(Y1,weights1,bias1,DataFormat="CB"); Y1 = reshape(Y1,numHiddenUnits,numFeatures,miniBatchSize); % Fully connect. Y2 = fullyconnect(hidden,weights2,bias2,DataFormat="CB"); Y2 = reshape(Y2,numHiddenUnits,1,miniBatchSize); % Addition, tanh. scores = tanh(Y1 + Y2); scores = reshape(scores, numHiddenUnits, numFeatures*miniBatchSize); % Fully connect, softmax. attentionWeights = fullyconnect(scores,weightsV,biasV,DataFormat="CB"); attentionWeights = reshape(attentionWeights,1,numFeatures,miniBatchSize); attentionWeights = softmax(attentionWeights,DataFormat="SCB"); % Context. contextVector = attentionWeights .* features; contextVector = squeeze(sum(contextVector,2)); end
Embedding Function
The embedding function maps an array of indices to a sequence of embedding vectors.
function Z = embedding(X, weights) % Reshape inputs into a vector [N, T] = size(X, 1:2); X = reshape(X, N*T, 1); % Index into embedding matrix Z = weights(:, X); % Reshape outputs by separating out batch and sequence dimensions Z = reshape(Z, [], N, T); end
Feature Extraction Function
The extractImageFeatures function takes as input a trained dlnetwork object, an input image, statistics for image rescaling, and the execution environment, and returns a dlarray containing the features extracted from the pretrained network.
function X = extractImageFeatures(net,X,inputMin,inputMax,executionEnvironment) % Resize and rescale. inputSize = net.Layers(1).InputSize(1:2); X = imresize(X,inputSize); X = rescale(X,-1,1,InputMin=inputMin,InputMax=inputMax); % Convert to dlarray. X = dlarray(X,"SSCB"); % Convert to gpuArray. if (executionEnvironment == "auto" && canUseGPU) || executionEnvironment == "gpu" X = gpuArray(X); end % Extract features and reshape. X = predict(net,X); sz = size(X); numFeatures = sz(1) * sz(2); inputSizeEncoder = sz(3); miniBatchSize = sz(4); X = reshape(X,[numFeatures inputSizeEncoder miniBatchSize]); end
Batch Creation Function
The createBatch function takes as input a mini-batch of data, tokenized captions, a pretrained network, statistics for image rescaling, a word encoding, and the execution environment, and returns a mini-batch of data corresponding to the extracted image features and captions for training.
function [X, T] = createBatch(X,documents,net,inputMin,inputMax,enc,executionEnvironment) X = extractImageFeatures(net,X,inputMin,inputMax,executionEnvironment); % Convert documents to sequences of word indices. T = doc2sequence(enc,documents,PaddingDirection="right",PaddingValue=enc.NumWords+1); T = cat(1,T{:}); % Convert mini-batch of data to dlarray. T = dlarray(T); % If training on a GPU, then convert data to gpuArray. if (executionEnvironment == "auto" && canUseGPU) || executionEnvironment == "gpu" T = gpuArray(T); end end
Encoder Model Function
The modelEncoder function takes as input an array of activations X and passes it through a fully connected operation and a ReLU operation. For the fully connected operation, operate on the channel dimension only. To apply the fully connected operation across the channel dimension only, flatten the other channels into a single dimension and specify this dimension as the batch dimension using the DataFormat option of the fullyconnect function.
function Y = modelEncoder(X,parametersEncoder) [numFeatures,inputSizeEncoder,miniBatchSize] = size(X); % Fully connect weights = parametersEncoder.fc.Weights; bias = parametersEncoder.fc.Bias; embeddingDimension = size(weights,1); X = permute(X,[2 1 3]); X = reshape(X,inputSizeEncoder,numFeatures*miniBatchSize); Y = fullyconnect(X,weights,bias,DataFormat="CB"); Y = reshape(Y,embeddingDimension,numFeatures,miniBatchSize); % ReLU Y = relu(Y); end
Decoder Model Function
The modelDecoder function takes as input a single time-step X, the decoder model parameters, the features from the encoder, and the network state, and returns the predictions for the next time step, the updated network state, and the attention weights.
function [Y,state,attentionWeights] = modelDecoder(X,parametersDecoder,features,state) hiddenState = state.gru.HiddenState; % Attention weights1 = parametersDecoder.attention.Weights1; bias1 = parametersDecoder.attention.Bias1; weights2 = parametersDecoder.attention.Weights2; bias2 = parametersDecoder.attention.Bias2; weightsV = parametersDecoder.attention.WeightsV; biasV = parametersDecoder.attention.BiasV; [contextVector, attentionWeights] = attention(hiddenState,features,weights1,bias1,weights2,bias2,weightsV,biasV); % Embedding weights = parametersDecoder.emb.Weights; X = embedding(X,weights); % Concatenate Y = cat(1,contextVector,X); % GRU inputWeights = parametersDecoder.gru.InputWeights; recurrentWeights = parametersDecoder.gru.RecurrentWeights; bias = parametersDecoder.gru.Bias; [Y, hiddenState] = gru(Y, hiddenState, inputWeights, recurrentWeights, bias, DataFormat="CBT"); % Update state state.gru.HiddenState = hiddenState; % Fully connect weights = parametersDecoder.fc1.Weights; bias = parametersDecoder.fc1.Bias; Y = fullyconnect(Y,weights,bias,DataFormat="CB"); % Fully connect weights = parametersDecoder.fc2.Weights; bias = parametersDecoder.fc2.Bias; Y = fullyconnect(Y,weights,bias,DataFormat="CB"); end
Model Loss
The modelLoss function takes as input the encoder and decoder parameters, the encoder features X, and the target caption T, and returns the loss, the gradients of the encoder and decoder parameters with respect to the loss, and the predictions.
function [loss,gradientsEncoder,gradientsDecoder,YPred] = ... modelLoss(parametersEncoder,parametersDecoder,X,T) miniBatchSize = size(X,3); sequenceLength = size(T,2) - 1; vocabSize = size(parametersDecoder.emb.Weights,2); % Model encoder features = modelEncoder(X,parametersEncoder); % Initialize state numHiddenUnits = size(parametersDecoder.attention.Weights1,1); state = struct; state.gru.HiddenState = dlarray(zeros([numHiddenUnits miniBatchSize],"single")); YPred = dlarray(zeros([vocabSize miniBatchSize sequenceLength],"like",X)); loss = dlarray(single(0)); padToken = vocabSize; for t = 1:sequenceLength decoderInput = T(:,t); YReal = T(:,t+1); [YPred(:,:,t),state] = modelDecoder(decoderInput,parametersDecoder,features,state); mask = YReal ~= padToken; loss = loss + sparseCrossEntropyAndSoftmax(YPred(:,:,t),YReal,mask); end % Calculate gradients [gradientsEncoder,gradientsDecoder] = dlgradient(loss, parametersEncoder,parametersDecoder); end
Sparse Cross Entropy and Softmax Loss Function
The sparseCrossEntropyAndSoftmax takes as input the predictions Y, corresponding targets T, and sequence padding mask, and applies the softmax functions and returns the cross-entropy loss.
function loss = sparseCrossEntropyAndSoftmax(Y, T, mask) miniBatchSize = size(Y, 2); % Softmax. Y = softmax(Y,DataFormat="CB"); % Find rows corresponding to the target words. idx = sub2ind(size(Y), T', 1:miniBatchSize); Y = Y(idx); % Bound away from zero. Y = max(Y, single(1e-8)); % Masked loss. loss = log(Y) .* mask'; loss = -sum(loss,"all") ./ miniBatchSize; end
Beam Search Function
The beamSearch function takes as input the image features X, a beam index, the parameters for the encoder and decoder networks, a word encoding, and a maximum sequence length, and returns the caption words for the image using the beam search algorithm.
function [words,attentionScores] = beamSearch(X,beamIndex,parametersEncoder,parametersDecoder, ... enc,maxNumWords) % Model dimensions numFeatures = size(X,1); numHiddenUnits = size(parametersDecoder.attention.Weights1,1); % Extract features features = modelEncoder(X,parametersEncoder); % Initialize state state = struct; state.gru.HiddenState = dlarray(zeros([numHiddenUnits 1],"like",X)); % Initialize candidates candidates = struct; candidates.State = state; candidates.Words = "<start>"; candidates.Score = 0; candidates.AttentionScores = dlarray(zeros([numFeatures maxNumWords],"like",X)); candidates.StopFlag = false; t = 0; % Loop over words while t < maxNumWords t = t + 1; candidatesNew = []; % Loop over candidates for i = 1:numel(candidates) % Stop generating when stop token is predicted if candidates(i).StopFlag continue end % Candidate details state = candidates(i).State; words = candidates(i).Words; score = candidates(i).Score; attentionScores = candidates(i).AttentionScores; % Predict next token decoderInput = word2ind(enc,words(end)); [YPred,state,attentionScores(:,t)] = modelDecoder(decoderInput,parametersDecoder,features,state); YPred = softmax(YPred,DataFormat="CB"); [scoresTop,idxTop] = maxk(extractdata(YPred),beamIndex); idxTop = gather(idxTop); % Loop over top predictions for j = 1:beamIndex candidate = struct; candidateWord = ind2word(enc,idxTop(j)); candidateScore = scoresTop(j); if candidateWord == "<stop>" candidate.StopFlag = true; attentionScores(:,t+1:end) = []; else candidate.StopFlag = false; end candidate.State = state; candidate.Words = [words candidateWord]; candidate.Score = score + log(candidateScore); candidate.AttentionScores = attentionScores; candidatesNew = [candidatesNew candidate]; end end % Get top candidates [~,idx] = maxk([candidatesNew.Score],beamIndex); candidates = candidatesNew(idx); % Stop predicting when all candidates have stop token if all([candidates.StopFlag]) break end end % Get top candidate words = candidates(1).Words(2:end-1); attentionScores = candidates(1).AttentionScores; end
Glorot Weight Initialization Function
The initializeGlorot function generates an array of weights according to Glorot initialization.
function weights = initializeGlorot(numOut, numIn) varWeights = sqrt( 6 / (numIn + numOut) ); weights = varWeights * (2 * rand([numOut, numIn], "single") - 1); end
See Also
word2ind (Text Analytics Toolbox) | tokenizedDocument (Text Analytics Toolbox) | wordEncoding (Text Analytics Toolbox) | dlarray | adamupdate | dlupdate | dlfeval | dlgradient | crossentropy | softmax | lstm | doc2sequence (Text Analytics Toolbox) | gru