proteinplot
Open Protein Plot window to investigate properties of amino acid sequence
Description
proteinplot opens the Protein Plot window that enables you to
analyze and compare properties of a single amino acid sequence. It displays smoothed line
plots of various properties such as the hydrophobicity of the amino acids in the
sequence.
proteinplot( opens the Protein Plot
window and loads SeqAA)SeqAA, an amino acid sequence, into the window.
You can analyze and compare properties of an amino acid sequence from the MATLAB® command line also by using the proteinpropplot function.
Examples
You can import a sequence into the Protein Plot window from the MATLAB command line.
Retrieve an amino acid sequence from the Protein Data Bank (PDB) database.
prion = getpdb("1HJM",SEQUENCEONLY=true);Load the amino acid sequence into the Protein Plot window.
proteinplot(prion)
The Protein Plot window opens, and the sequence appears in the Sequence text box.
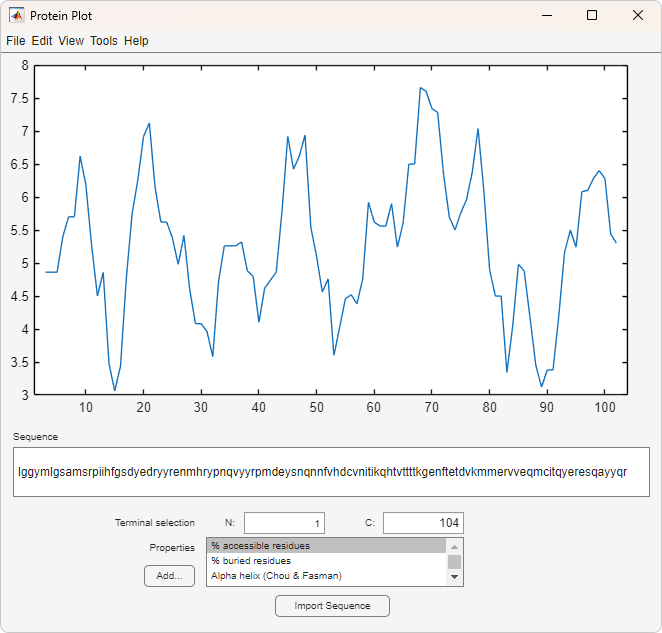
You can import a sequence after the Protein Plot window is open by doing either of the following:
Type or paste an amino acid sequence into the Sequence text box.
Click the Import Sequence button to open the Import dialog box From the Import From list, select one of the following:
Workspace — To select a variable from the MATLAB Workspace
Text File — To select a text file
FASTA File — To select a FASTA-formatted file
GenPept File — To select a GenPept-formatted file
GenPept Database — To specify an accession number in the GenPept database
Select a property from the Properties drop-down list box to display a smoothed plot of the property values along the sequence.
Select multiple properties from the list by holding down Shift or Ctrl while selecting properties.
When you select two properties, the plots are displayed using two y-axes, with one y-axis on the left and one on the right. For all other selections, a single
y-axis is displayed.When displaying one or two properties, the
yvalues displayed are the actual property values.When displaying three or more properties, the values are normalized to the range 0–1.
The Terminal Selection boxes (N and C) let you choose to plot only part of the sequence. By default, all of the sequence is plotted.
You can add your own properties by clicking on the Add button next to the Properties list. This opens a Property dialog box that lets you specify the value for each of the amino acids. The Display Text box lets you specify the text that will be displayed in the Properties list on the main Protein Plot window. You can also save the property values to a file for future use by typing a file name in the Filename text box.
The default smoothing method is an unweighted linear moving average with a window
length of five residues. You can change this by selecting Edit > Filter
Window Options. The dialog box lets you select the Window
Size from 5 to 29 residues.
Increasing the window size produces a smoother plot. You can modify the shape of the
smoothing window by changing the Edge Weight factor. And you can
choose the smoothing function to be a linear moving average, an exponential moving
average or a linear Lowess smoothing.
The File menu lets you import a sequence, save the plot that you have created to a Figure file, export the data values in the figure to a workspace variable or to a MAT-file, export the figure to a normal Figure window for customizing, or print the figure.
The Edit menu lets you create a new property, to reset the property values to the default values, and to modify the smoothing parameters with the Configuration Values menu item.
The View menu lets you turn the toolbar on and off, and to add a legend to the plot.
The Tools menu lets you zoom in and zoom out of the plot, to view Data Statistics such as mean, minimum and maximum values of the plot, and to normalize the values of the plot from
0to1.The Help menu lets you view this document and to see the references for the sequence properties included with the Protein Plot window.
Input Arguments
Amino acid sequence, specified as one of these values:
Character vector or string containing single-letter codes specifying an amino acid sequence. For valid letter codes, see the table Mapping Amino Acid Letter Codes to Integers. Unknown characters are mapped to
0.MATLAB structure containing a
Sequencefield that contains an amino acid sequence, such as returned byfastaread,getgenpept,genpeptread,getpdb, orpdbread.
Data Types: char | string | struct
Version History
Introduced before R2006a
See Also
aacount | atomiccomp | molweight | pdbdistplot | proteinpropplot | yyaxis
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)