ClassificationEnsemble
Ensemble classifier
Description
ClassificationEnsemble combines a set of
trained weak learner models and data on which these learners were trained. It can
predict ensemble response for new data by aggregating predictions from its weak
learners. It stores data used for training, can compute resubstitution predictions, and
can resume training if desired.
Creation
Description
Create a classification ensemble object (ens) using fitcensemble.
Properties
BinEdges — Bin edges for numeric predictors
cell array of p numeric vectors
This property is read-only.
Bin edges for numeric predictors, specified as a cell array of p numeric vectors, where p is the number of predictors. Each vector includes the bin edges for a numeric predictor. The element in the cell array for a categorical predictor is empty because the software does not bin categorical predictors.
The software bins numeric predictors only if you specify the 'NumBins'
name-value argument as a positive integer scalar when training a model with tree learners.
The BinEdges property is empty if the 'NumBins'
value is empty (default).
You can reproduce the binned predictor data Xbinned by using the
BinEdges property of the trained model
mdl.
X = mdl.X; % Predictor data
Xbinned = zeros(size(X));
edges = mdl.BinEdges;
% Find indices of binned predictors.
idxNumeric = find(~cellfun(@isempty,edges));
if iscolumn(idxNumeric)
idxNumeric = idxNumeric';
end
for j = idxNumeric
x = X(:,j);
% Convert x to array if x is a table.
if istable(x)
x = table2array(x);
end
% Group x into bins by using the discretize function.
xbinned = discretize(x,[-inf; edges{j}; inf]);
Xbinned(:,j) = xbinned;
endXbinned
contains the bin indices, ranging from 1 to the number of bins, for numeric predictors.
Xbinned values are 0 for categorical predictors. If
X contains NaNs, then the corresponding
Xbinned values are NaNs.
CategoricalPredictors — Indices of categorical predictors
vector of positive integers | []
This property is read-only.
Categorical predictor
indices, specified as a vector of positive integers. CategoricalPredictors
contains index values indicating that the corresponding predictors are categorical. The index
values are between 1 and p, where p is the number of
predictors used to train the model. If none of the predictors are categorical, then this
property is empty ([]).
Data Types: single | double
ClassNames — List of elements in Y with duplicates removed
categorical array | cell array of character vectors | character array | logical vector | numeric vector
This property is read-only.
List of the elements in Y with duplicates removed, returned as a
categorical array, cell array of character vectors, character array, logical vector, or
a numeric vector. ClassNames has the same data type as the data in
the argument Y. (The software treats string arrays as cell arrays of character
vectors.)
Data Types: double | logical | char | cell | categorical
CombineWeights — How the ensemble combines weak learner weights
'WeightedAverage' | 'WeightedSum'
This property is read-only.
How the ensemble combines weak learner weights, returned as either
'WeightedAverage' or 'WeightedSum'.
Data Types: char
Cost — Cost of classifying a point into class j when its true class is i
square matrix
Cost of classifying a point into class j when its true class is
i, returned as a square matrix. The rows of
Cost correspond to the true class and the columns correspond to
the predicted class. The order of the rows and columns of Cost
corresponds to the order of the classes in ClassNames. The number
of rows and columns in Cost is the number of unique classes in the
response.
Data Types: double
ExpandedPredictorNames — Expanded predictor names
cell array of character vectors
This property is read-only.
Expanded predictor names, returned as a cell array of character vectors.
If the model uses encoding for categorical variables, then
ExpandedPredictorNames includes the names that describe the
expanded variables. Otherwise, ExpandedPredictorNames is the same as
PredictorNames.
Data Types: cell
FitInfo — Fit information
numeric array
Fit information, returned as a numeric array. The FitInfoDescription property describes the content of this array.
Data Types: double
FitInfoDescription — Description of information in FitInfo
character vector | cell array of character vectors
Description of the information in FitInfo, returned as a character vector or cell array of character vectors.
Data Types: char | cell
HyperparameterOptimizationResults — Description of cross-validation optimization of hyperparameters
BayesianOptimization object | table of hyperparameters and associated values
This property is read-only.
Description of the cross-validation optimization of hyperparameters, returned as a
BayesianOptimization object or a table of
hyperparameters and associated values. Nonempty when the
OptimizeHyperparameters name-value pair is nonempty at creation.
Value depends on the setting of the HyperparameterOptimizationOptions
name-value pair at creation:
'bayesopt'(default) — Object of classBayesianOptimization'gridsearch'or'randomsearch'— Table of hyperparameters used, observed objective function values (cross-validation loss), and rank of observations from lowest (best) to highest (worst)
LearnerNames — Names of weak learners in ensemble
cell array of character vectors
This property is read-only.
Names of weak learners in ensemble, returned as a cell array of character vectors. The
name of each learner appears just once. For example, if you have an ensemble of 100
trees, LearnerNames is {'Tree'}.
Data Types: cell
Method — Method that creates ensemble
character vector
Method that fitcensemble uses to create the ensemble, returned as a character vector.
Example: 'AdaBoostM1'
Data Types: char
ModelParameters — Parameters used in training ensemble
EnsembleParams object
Parameters used in training the ensemble, returned as an EnsembleParams object. The properties of ModelParameters include the type of ensemble, either 'classification' or 'regression', the Method used to create the ensemble, and other parameters, depending on the ensemble.
NumObservations — Number of observations in the training data
positive integer
This property is read-only.
Number of observations in the training data, returned as a positive integer.
NumObservations can be less than the number of rows of input data
when there are missing values in the input data or response data.
Data Types: double
NumTrained — Number of trained weak learners
positive integer
This property is read-only.
Number of trained weak learners in the ensemble, returned as a positive integer.
Data Types: double
PredictorNames — Predictor names
cell array of character vectors
This property is read-only.
Predictor names, specified as a cell array of character vectors. The order of the
entries in PredictorNames is the same as in the training data.
Data Types: cell
Prior — Prior probabilities for each class
m-element vector
Prior probabilities for each class, returned as an m-element
vector, where m is the number of unique classes in the response. The
order of the elements of Prior corresponds to the order of the
classes in ClassNames.
Data Types: double
ReasonForTermination — Reason that fitcensemble stopped adding weak learners to ensemble
character vector
Reason that fitcensemble stopped adding weak learners to the ensemble, returned as a character vector.
Example: 'Terminated normally after completing the requested number of training cycles.'
Data Types: char
ResponseName — Name of the response variable
character vector
This property is read-only.
Name of the response variable, returned as a character vector.
Data Types: char
RowsUsed — Rows of the original predictor data X used for fitting
logical vector
This property is read-only.
Rows of the original predictor data X used for fitting, returned as an
n-element logical vector, where n is the
number of rows of X. If the software uses all rows of
X for constructing the object, then RowsUsed
is an empty array ([]).
Data Types: logical
ScoreTransform — Function for transforming scores
function handle | name of a built-in transformation function | 'none'
Function for transforming scores, specified as a function handle or the name of a built-in transformation function. 'none' means no transformation; equivalently, 'none' means @(x)x. For a list of built-in transformation functions and the syntax of custom transformation functions, see fitctree.
Add or change a ScoreTransform function using dot notation:
ctree.ScoreTransform = 'function' % or ctree.ScoreTransform = @function
Data Types: char | string | function_handle
Trained — Trained classification models
cell vector
Trained classification models, returned as a cell vector. The entries of the cell vector contain the corresponding compact classification models.
Data Types: cell
TrainedWeights — Trained weak learner weights
numeric vector
This property is read-only.
Trained weights for the weak learners in the ensemble, returned as a numeric vector.
TrainedWeights has T elements, where
T is the number of weak learners in
learners. The ensemble computes predicted response by aggregating
weighted predictions from its learners.
Data Types: double
UsePredForLearner — Indicator that learner j uses predictor i
logical matrix
Indicator that learner j uses predictor i,
returned as a logical matrix of size
P-by-NumTrained, where P is
the number of predictors (columns) in the training data.
UsePredForLearner(i,j) is true when learner
j uses predictor i, and is
false otherwise. For each learner, the predictors have the same
order as the columns in the training data.
If the ensemble is not of type Subspace, all entries in
UsePredForLearner are true.
Data Types: logical
W — Scaled weights in ensemble
numeric vector
This property is read-only.
Scaled weights in the ensemble, returned as a numeric vector. W has length n, the number of rows in the training data. The sum of the elements of W is 1.
Data Types: double
X — Predictor values
real matrix | table
This property is read-only.
Predictor values, returned as a real matrix or table. Each column of
X represents one variable (predictor), and each row represents
one observation.
Data Types: double | table
Y — Row classifications
categorical array | cell array of character vectors | character array | logical vector | numeric vector
This property is read-only.
Row classifications corresponding to the rows of X, returned as a categorical array, cell array of character vectors, character array, logical vector, or a numeric vector. Each row of Y represents the classification of the corresponding row of X.
Data Types: single | double | logical | char | string | cell | categorical
Object Functions
compact | Reduce size of classification ensemble model |
compareHoldout | Compare accuracies of two classification models using new data |
crossval | Cross-validate machine learning model |
edge | Classification edge for classification ensemble model |
gather | Gather properties of Statistics and Machine Learning Toolbox object from GPU |
lime | Local interpretable model-agnostic explanations (LIME) |
loss | Classification loss for classification ensemble model |
margin | Classification margins for classification ensemble model |
partialDependence | Compute partial dependence |
plotPartialDependence | Create partial dependence plot (PDP) and individual conditional expectation (ICE) plots |
predict | Predict labels using classification ensemble model |
predictorImportance | Estimates of predictor importance for classification ensemble of decision trees |
resubEdge | Resubstitution classification edge for classification ensemble model |
resubLoss | Resubstitution classification loss for classification ensemble model |
resubMargin | Resubstitution classification margins for classification ensemble model |
resubPredict | Classify observations in classification ensemble by resubstitution |
resume | Resume training of classification ensemble model |
shapley | Shapley values |
testckfold | Compare accuracies of two classification models by repeated cross-validation |
Examples
Train Boosted Classification Ensemble
Load the ionosphere data set.
load ionosphereTrain a boosted ensemble of 100 classification trees using all measurements and the AdaBoostM1 method.
Mdl = fitcensemble(X,Y,'Method','AdaBoostM1')
Mdl =
ClassificationEnsemble
ResponseName: 'Y'
CategoricalPredictors: []
ClassNames: {'b' 'g'}
ScoreTransform: 'none'
NumObservations: 351
NumTrained: 100
Method: 'AdaBoostM1'
LearnerNames: {'Tree'}
ReasonForTermination: 'Terminated normally after completing the requested number of training cycles.'
FitInfo: [100x1 double]
FitInfoDescription: {2x1 cell}
Mdl is a ClassificationEnsemble model object.
Mdl.Trained is the property that stores a 100-by-1 cell vector of the trained classification trees (CompactClassificationTree model objects) that compose the ensemble.
Plot a graph of the first trained classification tree.
view(Mdl.Trained{1},'Mode','graph')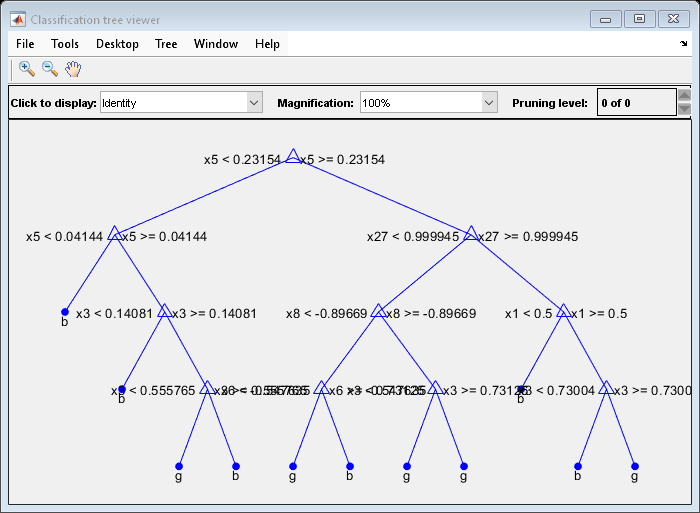
By default, fitcensemble grows shallow trees for boosted ensembles of trees.
Predict the label of the mean of X.
predMeanX = predict(Mdl,mean(X))
predMeanX = 1x1 cell array
{'g'}
Tips
For an ensemble of classification trees, the Trained property
of ens stores an ens.NumTrained-by-1
cell vector of compact classification models. For a textual or graphical
display of tree t in the cell vector, enter:
view(ens.Trained{for ensembles aggregated using LogitBoost or GentleBoost.t}.CompactRegressionLearner)view(ens.Trained{for all other aggregation methods.t})
Extended Capabilities
C/C++ Code Generation
Generate C and C++ code using MATLAB® Coder™.
Usage notes and limitations:
The
predictfunction supports code generation.To integrate the prediction of an ensemble into Simulink®, you can use the ClassificationEnsemble Predict block in the Statistics and Machine Learning Toolbox™ library or a MATLAB® Function block with the
predictfunction.When you train an ensemble by using
fitcensemble, the following restrictions apply.The value of the
ScoreTransformname-value argument cannot be an anonymous function.Code generation limitations for the weak learners used in the ensemble also apply to the ensemble.
For decision tree weak learners, you cannot use surrogate splits; that is, the value of the
Surrogatename-value argument must be'off'.For k-nearest neighbor weak learners, the value of the
Distancename-value argument cannot be a custom distance function. The value of theDistanceWeightname-value argument can be a custom distance weight function, but it cannot be an anonymous function.
For fixed-point code generation, the following additional restrictions apply.
When you train an ensemble by using
fitcensemble, you must train an ensemble using tree learners, and theScoreTransformvalue cannot be'invlogit'.Categorical predictors (
logical,categorical,char,string, orcell) are not supported. You cannot use theCategoricalPredictorsname-value argument. To include categorical predictors in a model, preprocess them by usingdummyvarbefore fitting the model.Class labels with the
categoricaldata type are not supported. Both the class label value in the training data (TblorY) and the value of theClassNamesname-value argument cannot be an array with thecategoricaldata type.
For more information, see Introduction to Code Generation.
GPU Arrays
Accelerate code by running on a graphics processing unit (GPU) using Parallel Computing Toolbox™.
Usage notes and limitations:
The following object functions fully support GPU arrays:
The following object functions offer limited support for GPU arrays:
The object functions execute on a GPU if at least one of the following applies:
The model was fitted with GPU arrays.
The predictor data that you pass to the object function is a GPU array.
For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2011aR2023b: Model with discriminant analysis weak learners stores observations with missing predictor values
Starting in R2023b, training observations with missing predictor values are
included in the X, Y, and
W data properties of classification ensemble models with
discriminant analysis weak learners. The RowsUsed property
indicates the training observations stored in the model, rather than those used for
training. Observations with missing predictor values continue to be omitted from the
model training process.
In previous releases, the software omitted training observations that contained missing predictor values from the data properties of the model.
R2022a: Cost property stores the user-specified cost matrix
Starting in R2022a, the Cost property stores the user-specified cost
matrix, so that you can compute the observed misclassification cost using the specified cost
value. The software stores normalized prior probabilities (Prior)
and observation weights (W) that do not reflect the penalties described
in the cost matrix. To compute the observed misclassification cost, specify the
LossFun name-value argument as "classifcost"
when you call the loss or resubLoss
function.
Note that model training has not changed and, therefore, the decision boundaries between classes have not changed.
For training, the fitting function updates the specified prior probabilities by
incorporating the penalties described in the specified cost matrix, and then normalizes the
prior probabilities and observation weights. This behavior has not changed. In previous
releases, the software stored the default cost matrix in the Cost
property and stored the prior probabilities and observation weights used for training in the
Prior and W properties, respectively. Starting
in R2022a, the software stores the user-specified cost matrix without modification, and stores normalized
prior probabilities and observation weights that do not reflect the cost penalties. For more
details, see Misclassification Cost Matrix, Prior Probabilities, and Observation Weights.
Some object functions use the Cost, Prior, and W properties:
The
lossandresubLossfunctions use the cost matrix stored in theCostproperty if you specify theLossFunname-value argument as"classifcost"or"mincost".The
lossandedgefunctions use the prior probabilities stored in thePriorproperty to normalize the observation weights of the input data.The
resubLossandresubEdgefunctions use the observation weights stored in theWproperty.
If you specify a nondefault cost matrix when you train a classification model, the object functions return a different value compared to previous releases.
If you want the software to handle the cost matrix, prior
probabilities, and observation weights in the same way as in previous releases, adjust the prior
probabilities and observation weights for the nondefault cost matrix, as described in Adjust Prior Probabilities and Observation Weights for Misclassification Cost Matrix. Then, when you train a
classification model, specify the adjusted prior probabilities and observation weights by using
the Prior and Weights name-value arguments, respectively,
and use the default cost matrix.
See Also
ClassificationTree | fitcensemble | CompactClassificationEnsemble | view | compareHoldout
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.

Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list:
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)