movmean
Moving mean
Syntax
Description
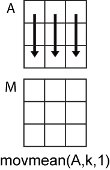
M = movmean( returns the
local A,k)k-point mean values, where each
mean is calculated over a sliding window of length k across
neighboring elements of A. When k is odd, the
window is centered about the element in the current position. When
k is even, the window is centered about the current and
previous elements. The window size is automatically truncated at the endpoints when
there are not enough elements to fill the window. When the window is truncated, the
average is taken over only the elements that fill the window. M
is the same size as A.
If
Ais a vector, thenmovmeanoperates along the length of the vectorA.If
Ais a multidimensional array, thenmovmeanoperates along the first dimension ofAwhose size does not equal 1.If
Ais a table or timetable, thenmovmeanoperates along the variables ofA. (since R2025a)
M = movmean(___, specifies the
dimension of dim)A to operate along for any of the previous syntaxes.
For example, if A is a matrix, then
movmean(A,k,2) operates along the columns of
A, computing the k-element sliding mean
for each row.
M = movmean(___, specifies
whether to include or omit nanflag)NaN values in A.
For example, movmean(A,k,"omitnan") ignores
NaN values when computing each mean. By default,
movmean includes NaN values.
M = movmean(___, specifies
additional parameters for the moving average using one or more name-value arguments.
For example, if Name,Value)x is a vector of time values, then
movmean(A,k,"SamplePoints",x) computes the moving average
relative to the times in x.
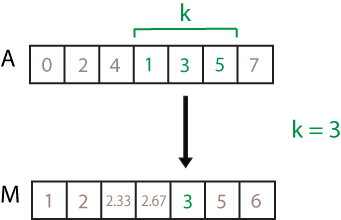
![movmean(A,[2 1]) computation. The elements in the sample window are 4, 1, 3, and 5, so the resulting local mean is 3.25.](movmean_windowing.png)