makima
Modified Akima piecewise cubic Hermite interpolation
Description
Examples
Use makima to interpolate a cosine curve over unevenly spaced sample points.
x = [0 1 2.5 3.6 5 7 8.1 10]; y = cos(x); xq = 0:.25:10; yq = makima(x,y,xq); plot(x,y,'o',xq,yq,'--')
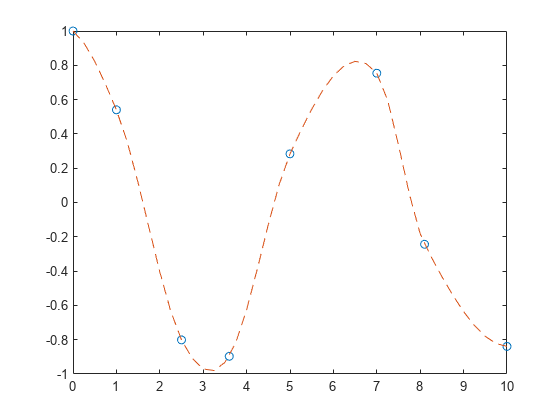
With oscillatory functions, the Akima algorithm flattens the curve near local extrema. To compensate for this flattening, you can add more sample points near the local extrema.
Add sample points near and and replot the interpolation.
x = [0 1 2.5 3.6 5 6.5 7 8.1 9 10]; y = cos(x); xq = 0:.25:10; yq = makima(x,y,xq); plot(x,y,'o',xq,yq,'--')
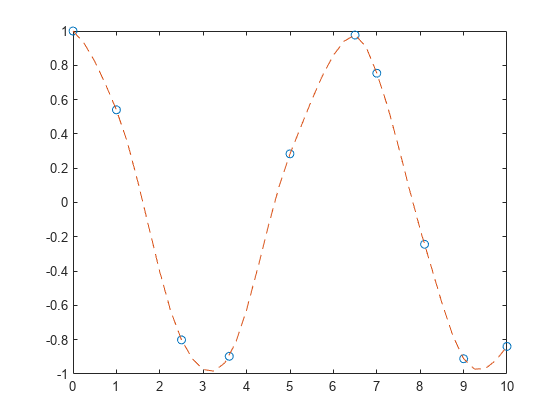
Compare the interpolation results produced by spline, pchip, and makima for two different data sets. These functions all perform different forms of piecewise cubic Hermite interpolation. Each function differs in how it computes the slopes of the interpolant, leading to different behaviors when the underlying data has flat areas or undulations.
Compare the interpolation results on sample data that connects flat regions. Create vectors of x values, function values at those points y, and query points xq. Compute interpolations at the query points using spline, pchip, and makima. Plot the interpolated function values at the query points for comparison.
x = -3:3; y = [-1 -1 -1 0 1 1 1]; xq1 = -3:.01:3; p = pchip(x,y,xq1); s = spline(x,y,xq1); m = makima(x,y,xq1); plot(x,y,'o',xq1,p,'-',xq1,s,'-.',xq1,m,'--') legend('Sample Points','pchip','spline','makima','Location','SouthEast')

In this case, pchip and makima have similar behavior in that they avoid overshoots and can accurately connect the flat regions.
Perform a second comparison using an oscillatory sample function.
x = 0:15; y = besselj(1,x); xq2 = 0:0.01:15; p = pchip(x,y,xq2); s = spline(x,y,xq2); m = makima(x,y,xq2); plot(x,y,'o',xq2,p,'-',xq2,s,'-.',xq2,m,'--') legend('Sample Points','pchip','spline','makima')

When the underlying function is oscillatory, spline and makima capture the movement between points better than pchip, which is aggressively flattened near local extrema.
Create vectors for the sample points x and values at those points y. Use makima to construct a piecewise polynomial structure for the data.
x = -5:5; y = [1 1 1 0 0 1 1 2 2 2 2]; pp = makima(x,y)
pp = struct with fields:
form: 'pp'
breaks: [-5 -4 -3 -2 -1 0 1 2 3 4 5]
coefs: [10×4 double]
pieces: 10
order: 4
dim: 1
The structure contains the information for 10 polynomials of order 4 that span the data. pp.coefs(i,:) contains the coefficients for the polynomial that is valid in the region defined by the breakpoints [breaks(i) breaks(i+1)].
Use the structure with ppval to evaluate the interpolation at several query points, and then plot the results. In regions with three or more constant points, the Akima algorithm connects the points with a straight line.
xq = -5:0.2:5; m = ppval(pp,xq); plot(x,y,'o',xq,m,'-.') ylim([-0.2 2.2])
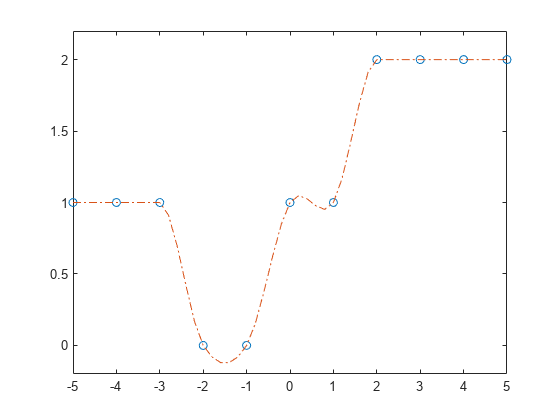
Input Arguments
Sample points, specified as a vector. The vector x specifies the
points at which the data y is given. The elements of
x must be unique.
Data Types: single | double
Function values at sample points, specified as a numeric vector, matrix, or array.
x and y must have the same length.
If y is a matrix or array, then the values in the last dimension,
y(:,...,:,j), are taken as the values to match with
x. In that case, the last dimension of y must be
the same length as x.
Data Types: single | double
Query points, specified as a scalar, vector, matrix, or array. The points specified
in xq are the x-coordinates for the interpolated
function values yq computed by makima.
Data Types: single | double
Output Arguments
Interpolated values at query points, returned as a scalar, vector, matrix, or array.
The size of yq is related to the sizes of y and
xq:
If
yis a vector, thenyqhas the same size asxq.If
yis an array of sizeNy = size(y), then these conditions apply:If
xqis a scalar or vector, thensize(yq)returns[Ny(1:end-1) length(xq)].If
xqis an array, thensize(yq)returns[Ny(1:end-1) size(xq)].
Piecewise polynomial, returned as a structure. Use this structure with the ppval function to evaluate the interpolating polynomials at one or more
query points. The structure has these fields.
| Field | Description |
|---|---|
form |
|
breaks | Vector of length |
coefs |
|
pieces | Number of pieces, |
order | Order of polynomials |
dim | Dimensionality of target |
Because the polynomial coefficients in coefs are local
coefficients for each interval, you must subtract the lower endpoint of the
corresponding knot interval to use the coefficients in a conventional polynomial
equation. In other words, for the coefficients [a,b,c,d] on the
interval [x1,x2], the corresponding polynomial is
More About
The Akima algorithm for one-dimensional interpolation, described in [1] and [2], performs cubic interpolation to produce piecewise polynomials with continuous first-order derivatives (C1). The algorithm avoids excessive local undulations.
If is the slope on interval , then the value of the derivative at the sample point is a weighted average of nearby slopes:
In Akima's original formula, the weights are:
The original Akima algorithm gives equal weight to the points on both sides, evenly dividing an undulation.
When two flat regions with different slopes meet, the modification made to the original Akima algorithm gives more weight to the side where the slope is closer to zero. This modification gives priority to the side that is closer to horizontal, which is more intuitive and avoids overshoot. In particular, whenever there are three or more consecutive collinear points, the algorithm connects them with a straight line and thus avoids an overshoot.
The weights used in the modified Akima algorithm are:
Compared to the spline algorithm, the Akima algorithm produces
fewer undulations and is better suited to deal with quick changes between flat regions.
Compared to the pchip algorithm, the Akima algorithm is not as
aggressively flattened and is therefore still able to deal with oscillatory data.
References
[1] Akima, Hiroshi. "A new method of interpolation and smooth curve fitting based on local procedures." Journal of the ACM (JACM) , 17.4, 1970, pp. 589–602.
[2] Akima, Hiroshi. "A method of bivariate interpolation and smooth surface fitting based on local procedures." Communications of the ACM , 17.1, 1974, pp. 18–20.
Extended Capabilities
Usage notes and limitations:
Input
xmust be strictly increasing.Code generation does not remove
yentries withNaNvalues.If you generate code for the
pp = makima(x,y)syntax, then you cannot inputppto theppvalfunction in MATLAB®. To create a MATLABppstructure from appstructure created by the code generator:In code generation, use
unmkppto return the piecewise polynomial details to MATLAB.In MATLAB, use
mkppto create theppstructure.
If you supply
xq, and ifyhas a variable-size and is not a variable-length vector, then the orientation of vector outputs in the generated code might not match the orientation in MATLAB.
Refer to the usage notes and limitations in the C/C++ Code Generation section. The same usage notes and limitations apply to GPU code generation.
The makima function fully supports
thread-based environments. For more information, see Run MATLAB Functions in Thread-Based Environment.
The makima function
fully supports GPU arrays. To run the function on a GPU, specify the input data as a gpuArray (Parallel Computing Toolbox). For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
The makima function
supports distributed arrays with these usage notes and limitations:
The
makima(syntax is not supported for distributed arrays.x,y)
For more information, see Run MATLAB Functions with Distributed Arrays (Parallel Computing Toolbox).
Version History
Introduced in R2019b
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)