Preprocess Multiresolution Images for Training Classification Network
This example shows how to prepare datastores that read and preprocess multiresolution whole slide images (WSIs) that might not fit in memory.
Deep learning approaches to tumor classification rely on digital pathology, in which whole tissue slides are imaged and digitized. The resulting WSIs have high resolution, on the order of 200,000-by-100,000 pixels. WSIs are frequently stored in a multiresolution format to facilitate efficient display, navigation, and processing of images.
Read and process WSI data using blockedImage and blockedImageDatastore objects. These objects facilitate working with multiple resolution levels and do not require the image to be loaded into core memory. This example shows how to use lower resolution image data to efficiently prepare data from the finer levels. You can use the processed data to train classification deep learning networks. For an example, see Classify Tumors in Multiresolution Blocked Images.
Download Camelyon16 Data Set
This example uses WSIs from the Camelyon16 challenge [1]. The data from this challenge contains a total of 400 WSIs of lymph nodes from two independent sources, separated into 270 training images and 130 test images. The WSIs are stored as TIF files in a stripped format with an 11-level pyramid structure.
The training data set consists of 159 WSIs of normal lymph nodes and 111 WSIs of lymph nodes with tumor and healthy tissue. Ground truth coordinates of the lesion boundaries accompany the tumor images. The size of the data set is approximately 451 GB.
First, create folders where you want to store the downloaded data on your machine.
dataDir = fullfile(tempdir,"Camelyon16"); trainingDir = fullfile(dataDir,"training"); trainNormalDir = fullfile(trainingDir,"normal"); trainTumorDir = fullfile(trainingDir,"tumor"); trainAnnotationDir = fullfile(trainingDir,"lesion_annotations"); if ~exist(dataDir,"dir") mkdir(trainingDir) mkdir(trainNormalDir) mkdir(trainTumorDir) mkdir(trainAnnotationDir) end
To download the data, go to the GigaDB website for the data set, and, in the table, navigate to the Files tab. Download individual files by clicking on the name in the File Name column, and move them to the correct subfolders by following these steps:
Download all TIF images with filenames starting with
normal_, and store them in the folder specified by thetrainNormalDirvariable.Download all TIF images with filenames starting with
tumor_, and store them in the folder specified by thetrainTumorDirvariable.Download the
lesion_annotations.zipfile for the CAMELYON16 training data set, and extract the zipped files to the folder specified by thetrainAnnotationDirvariable. You can differentiate between the lesions file for the training and testing data set using the information in the Description column of the table.
Create blockedImage Objects to Manage WSI Images
Get the file names of the normal and tumor training images.
normalFileSet = matlab.io.datastore.FileSet(fullfile(trainNormalDir,"normal*")); normalFilenames = normalFileSet.FileInfo.Filename; tumorFileSet = matlab.io.datastore.FileSet(fullfile(trainTumorDir,"tumor*")); tumorFilenames = tumorFileSet.FileInfo.Filename;
One of the training images of normal tissue, "normal_144.tif," is missing important metadata required to deduce the physical extents of the image. Exclude this file.
normalFilenames(contains(normalFilenames,"normal_144")) = [];Create two arrays of blockedImage objects, one for normal images and one for the tumor images. Each blockedImage object points to the corresponding image file on disk.
normalImages = blockedImage(normalFilenames); tumorImages = blockedImage(tumorFilenames);
Display WSI Images
To get a better understanding of the training data, display the blocked images of normal tissue. The images at a coarse resolution level are small enough to fit in memory, so you can visually inspect the blockedImage data in the Image Browser app using the browseBlockedImages helper function. This helper function is attached to the example as a supporting file. Note that the images contain a lot of empty white space and that the tissue occupies only a small portion of the image. These characteristics are typical of WSIs.
browseBlockedImages(normalImages,8)

Explore a single blockedImage in more depth. Select a sample tumor image to visualize, then display the image using the imageshow function. The function automatically selects the resolution level based on the screen size and the current zoom level.
idxSampleTumorImage = 10; tumorImage = tumorImages(idxSampleTumorImage); h = imageshow(tumorImage);
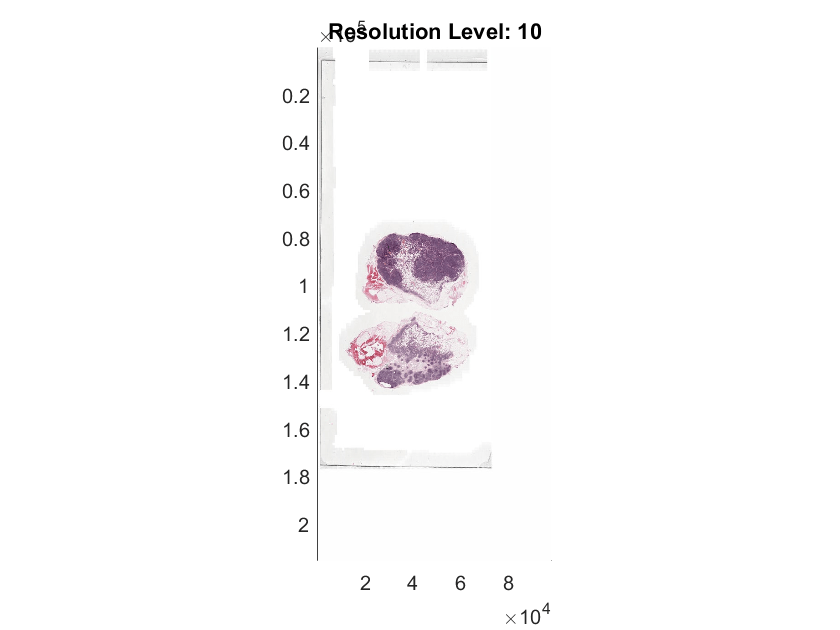
h.Parent.Title = "Resolution Level: "+h.ResolutionLevel;Zoom in to an area of interest. The resolution level automatically increases to show more detail. Extract the region using the locations of top-left and bottom-right corners of the ROI.
topLocation = [33600 104600]; bottomLocation = [35000 106000]; roiStart = [topLocation(2),topLocation(1)]; roiEnd = [bottomLocation(2),bottomLocation(1)]; imRegion = getRegion(tumorImage,roiStart,roiEnd); he = imageshow(imRegion);

he.Parent.Title = "Resolution Level: "+he.ResolutionLevel;Align Spatial Extents
When you work with multiple resolution levels of a multiresolution image, the spatial extents of each level must match. If spatial extents are aligned, then information such as masks can be extracted at coarse resolution levels and correctly applied to matching locations at finer resolution levels. For more information, see Set Up Spatial Referencing for Blocked Images.
Inspect the dimensions of the sample tumor image at each resolution level. Level 1 has the most pixels and is the finest resolution level. Level 10 has the fewest pixels and is the coarsest resolution level. The aspect ratio is not consistent, which usually indicates that the levels do not span the same world area.
levelSizeInfo = table((1:length(tumorImage.Size))', ... tumorImage.Size(:,1), ... tumorImage.Size(:,2), ... tumorImage.Size(:,1)./tumorImage.Size(:,2), ... VariableNames=["Resolution Level" "Image Width" "Image Height" "Aspect Ratio"]); disp(levelSizeInfo)
Resolution Level Image Width Image Height Aspect Ratio
________________ ___________ ____________ ____________
1 2.1555e+05 97792 2.2042
2 1.0803e+05 49152 2.1979
3 54272 24576 2.2083
4 27136 12288 2.2083
5 13824 6144 2.25
6 7168 3072 2.3333
7 1577 3629 0.43455
8 3584 1536 2.3333
9 2048 1024 2
10 1024 512 2
11 512 512 1
Set the spatial referencing for all training data by using the setSpatialReferencingForCamelyon16 helper function. This function is attached to the example as a supporting file. The setSpatialReferencingForCamelyon16 function sets the WorldStart and WorldEnd properties of each blockedImage object using the spatial referencing information from the TIF file metadata.
normalImages = setSpatialReferencingForCamelyon16(normalImages); tumorImages = setSpatialReferencingForCamelyon16(tumorImages);
Create Tissue Masks
The majority of a typical WSI consists of background pixels. To process WSI data efficiently, you can create a region of interest (ROI) mask from a coarse resolution level, and then limit computations at finer resolution levels to regions within the ROI. For more information, see Process Blocked Images Efficiently Using Mask.
Consider these two factors when picking a mask level:
The image size at the chosen mask level. To process the masks more quickly, use a lower resolution level.
The fidelity of ROI boundaries at the chosen level. To capture the boundary accurately, use a higher resolution level.
This example uses a coarse resolution level to create masks for the training images of normal tissue.
normalMaskLevel = 8;
The background is uniformly light. You can segment out the tissue through a simple thresholding operation. Apply a threshold in a block-wise manner to all normal training images using the apply function. Save the results to disk by specifying the OutputLocation name-value argument.
trainNormalMaskDir = fullfile(trainingDir,"normal_mask_level"+num2str(normalMaskLevel)); if ~isfolder(trainNormalMaskDir) normalMasks = apply(normalImages, @(bs)imclose(im2gray(bs.Data)<150, ones(5)), ... BlockSize=[512 512],... Level=normalMaskLevel, ... UseParallel=canUseGPU, ... OutputLocation=trainNormalMaskDir); save(fullfile(trainNormalMaskDir,"normalMasks.mat"),"normalMasks") else load(fullfile(trainNormalMaskDir,"normalMasks.mat"),"normalMasks"); end
Starting parallel pool (parpool) using the 'Processes' profile ... Connected to parallel pool with 12 workers.
Display Tissue Masks
The tissue masks have only one level and are small enough to fit in memory. Display the tissue masks in the Image Browser app using the browseBlockedImages helper function. This helper function is attached to the example as a supporting file.
browseBlockedImages(normalMasks,1)

Create Tumor Masks
In tumor images, the ROI consists of tumor tissue. The color of tumor tissue is similar to the color of healthy tissue, so you cannot use color segmentation techniques. Instead, create ROIs by using the ground truth coordinates of the lesion boundaries that accompany the tumor images. These regions are hand drawn at the finest level.
Display Tumor Boundaries
To get a better understanding of the tumor training data, read the ground truth boundary coordinates and display the coordinates as freehand ROIs using the showCamelyon16TumorAnnotations helper function. This helper function is attached to the example as a supporting file. Notice that normal regions (shown with a green boundary) can occur inside tumor regions. Zoom in to inspect an area of interest.
idxSampleTumorImage = 11; tumorImage = tumorImages(idxSampleTumorImage); showCamelyon16TumorAnnotations(tumorImage,trainAnnotationDir);
Convert Polygon Coordinates to Binary Blocked Images
Specify the resolution level of the tumor masks.
tumorMaskLevel = 8;
Create a tumor mask for each image using the createMaskForCamelyon16TumorTissue helper function. This helper function is attached to the example as a supporting file. For each image, the function performs these operations.
Read the (x, y) boundary coordinates for all ROIs in the annotated XML file.
Separate the boundary coordinates for tumor and normal tissue ROIs into separate cell arrays.
Convert the cell arrays of boundary coordinates to a binary blocked image using the
polyToBlockedImagefunction. In the binary image, the ROI indicates tumor pixels and the background indicates normal tissue pixels. Pixels that are within both tumor and normal tissue ROIs are classified as background.
trainTumorMaskDir = fullfile(trainingDir,"tumor_mask_level"+num2str(tumorMaskLevel)); if ~isfolder(trainTumorMaskDir) mkdir(trainTumorMaskDir) tumorMasks = createMaskForCamelyon16TumorTissue( ... tumorImages,trainAnnotationDir,trainTumorMaskDir,tumorMaskLevel); save(fullfile(trainTumorMaskDir,"tumorMasks.mat"),"tumorMasks") else load(fullfile(trainTumorMaskDir,"tumorMasks.mat"),"tumorMasks"); end
The tumor masks have only one resolution level and are small enough to fit in memory. Display the tumor masks in the Image Browser app using the browseBlockedImages helper function. This helper function is attached to the example as a supporting file.
browseBlockedImages(tumorMasks,1)
Confirm Fidelity of Mask to ROI Boundaries
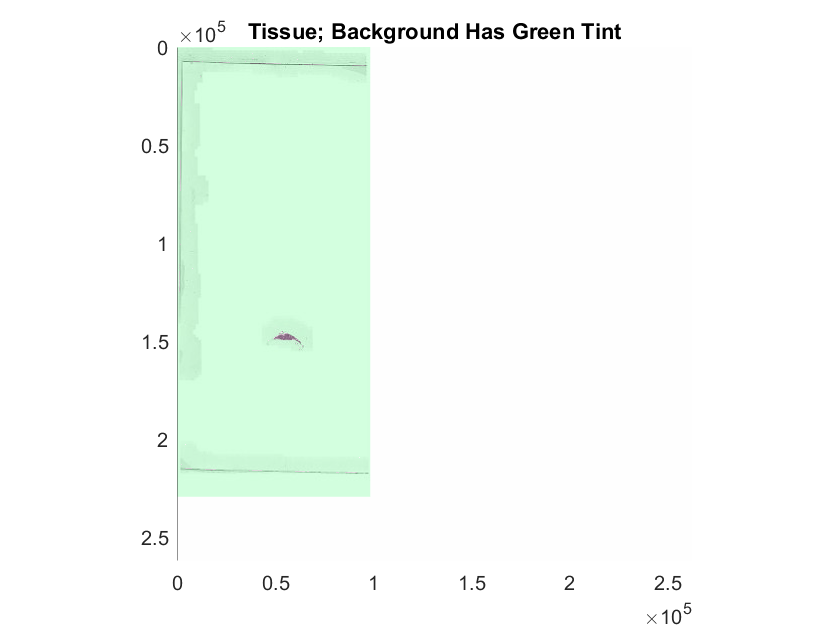
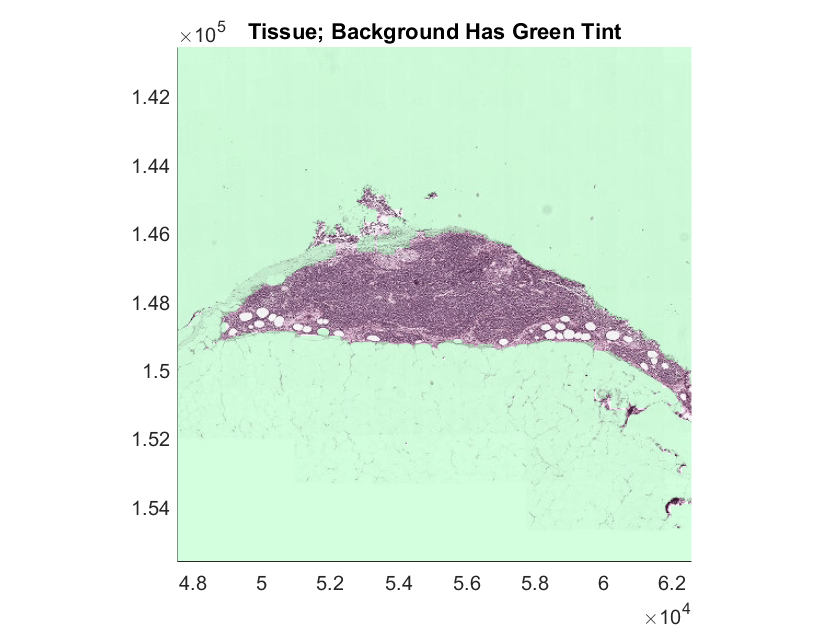
Select a sample tumor image that has intricate regions, and display the blockedImage using the imageshow function. Display the mask as an overlay on the blockedImage. Set the transparency of each pixel using the OverlayAlphamap name-value argument. For pixels within the ROI, the labels are fully transparent. For pixels outside the ROI, the label is partially transparent and appears with a green tint.
idxSampleTumorImage = 11;
tumorImage = tumorImages(idxSampleTumorImage);
tumorMask = tumorMasks(idxSampleTumorImage);
tumorImg = gather(tumorImage,Level=8);
hTumor = imageshow(tumorImg,OverlayData=tumorMask,OverlayDisplayRangeMode="data-range",OverlayColormap=[0 1 0],OverlayAlphamap=[0.3 0]);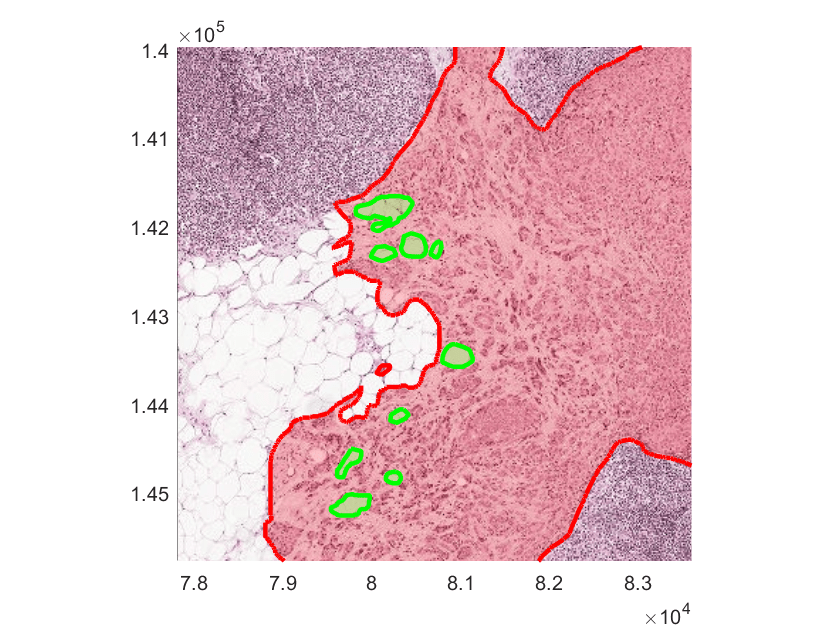
hTumor.Parent.Title = "Tumor; Background Has Green Tint";Select Blocks for Training
You can train a network using data from any resolution level. Finer resolution levels provide more homogenous blocks for either class. Coarser levels, which cover a larger spatial region for the same block size, have more surrounding context. For this example, select blocks at the finest resolution level.
trainingLevel = 1;
Specify the block size to match the input size of the network. This example uses a block size suitable for the Inception-v3 network.
networkBlockSize = [299 299 3];
Create sets of normal and tumor blocks using the selectBlockLocations function. This function selects blocks that are within the specified mask area. You can refine the number of selected blocks for each class by specifying the BlockOffsets and InclusionThreshold name-value arguments. Consider these two factors when calling the selectBlockLocations function.
The amount of training data. Using as much of the training data as possible helps to generalize the network during training and ensures a good class representation balance in the selected block set. Increase the number of selected blocks by decreasing the
BlockOffsetsandInclusionThresholdname-value arguments.The hardware and time available to train. Using more blocks requires more training time or more powerful hardware. Decrease the number of selected blocks by increasing the
BlockOffsetsandInclusionThresholdname-value arguments.
Select blocks within the normal images using the tissue masks. This example specifies values of BlockOffsets and InclusionThreshold that result in a relatively small number of blocks.
normalOffsetFactor = 3.5; normalInclusionThreshold = 0.97; blsNormalData = selectBlockLocations(normalImages, ... BlockSize=networkBlockSize(1:2), ... BlockOffsets=round(networkBlockSize(1:2)*normalOffsetFactor), ... Levels=trainingLevel, ... Masks=normalMasks, ... InclusionThreshold=normalInclusionThreshold, ... ExcludeIncompleteBlocks=true, ... UseParallel=canUseGPU); disp(blsNormalData)
blockLocationSet with properties:
ImageNumber: [4198×1 double]
BlockOrigin: [4198×3 double]
BlockSize: [299 299 3]
Levels: [1 1 1 1 1 1 1 1 1 1]
Select blocks within the tumor images using the tumor masks. This example specifies values of BlockOffsets and InclusionThreshold that result in a relatively small number of blocks.
tumorOffsetFactor = 1; tumorInclusionThreshold = 0.90; blsTumorData = selectBlockLocations(tumorImages, ... BlockSize=networkBlockSize(1:2), ... BlockOffsets=round(networkBlockSize(1:2)*tumorOffsetFactor), ... Levels=trainingLevel, ... Masks=tumorMasks, ... InclusionThreshold=tumorInclusionThreshold, ... ExcludeIncompleteBlocks=true, ... UseParallel=canUseGPU); disp(blsTumorData)
blockLocationSet with properties:
ImageNumber: [10898×1 double]
BlockOrigin: [10898×3 double]
BlockSize: [299 299 3]
Levels: [1 1 1 1 1 1 1 1 1 1 1]
Create Blocked Image Datastores for Training and Validation
Create a single blockedImageDatastore object by combining the sets of normal and tumor blocks.
[blsAll,allImages] = mergeBlockLocationSets(blsNormalData,normalImages,blsTumorData,tumorImages); dsAllBlocks = blockedImageDatastore(allImages,BlockLocationSet=blsAll);
Shuffle the blocks to reduce chances of overfitting and to increase generalization in the training process.
dsAllBlocks = shuffle(dsAllBlocks);
Partition the blocks into training and validation data sets. Allocate 99% of the blocks for training and use the remaining 1% for validation.
numericBlockLabels = [zeros(size(blsNormalData.ImageNumber)); ones(size(blsTumorData.ImageNumber))]; blockLabels = categorical(numericBlockLabels,[0,1],["normal","tumor"]); idx = splitlabels(blockLabels,0.99,"randomized"); dsTrain = subset(dsAllBlocks,idx{1}); dsVal = subset(dsAllBlocks,idx{2});
Training a classification network requires labeled training data. Label each block as normal or tumor based on the image containing the block. Specify the block labels as normal and tumor.
numericImageLabels = [zeros(size(normalImages)),ones(size(tumorImages))]; imageLabels = categorical(numericImageLabels,[0,1],["normal","tumor"]);
Transform the blockedImageDatastore such that the datastore returns both blocks and corresponding labels using the transform function and the labelCamelyon16Blocks helper function. The helper function is attached to the example as a supporting file.
The labelCamelyon16Blocks helper function derives the block label from the image index, which is stored in the ImageNumber field of the block metadata. The helper function returns the block data and label as a two-element cell array, suitable for training a classification network.
dsTrainLabeled = transform(dsTrain, ... @(block,blockInfo) labelCamelyon16Blocks(block,blockInfo,imageLabels),IncludeInfo=true); dsValLabeled = transform(dsVal, ... @(block,blockInfo) labelCamelyon16Blocks(block,blockInfo,imageLabels),IncludeInfo=true);
Augment Training Data
Augment the training data using the transform function and the augmentBlocksReflectionRotation helper function. The helper function is attached to the example as a supporting file.
The augmentBlocksReflectionRotation helper function increases the size of the training data by creating three variations of each input block with reflections and 90 degree rotations.
dsTrainLabeled = transform(dsTrainLabeled,@augmentBlocksReflectionRotation);
Preview a batch of training data.
batch = preview(dsTrainLabeled);
h = uifigure;
g = uigridlayout(h,ColumnWidth={"1x","1x","1x","1x"},RowHeight={"fit","1x","1x","1x"});
viewer1 = viewer2d(g,ScaleBar="off");
viewer2 = viewer2d(g,ScaleBar="off");
viewer3 = viewer2d(g,ScaleBar="off");
viewer4 = viewer2d(g,ScaleBar="off");
viewer1.Layout.Row = 1;
viewer1.Layout.Column = 1;
viewer2.Layout.Row = 1;
viewer2.Layout.Column = 2;
viewer3.Layout.Row = 1;
viewer3.Layout.Column = 3;
viewer4.Layout.Row = 1;
viewer4.Layout.Column = 4;
imageshow(batch{1,1},Parent=viewer1);
imageshow(batch{2,1},Parent=viewer2);
imageshow(batch{3,1},Parent=viewer3);
imageshow(batch{4,1},Parent=viewer4); 
You can improve the ability of the network to generalize to other data by performing additional randomized augmentation operations. For example, stain normalization is a common augmentation technique for WSI images. Stain normalization reduces the variability in color and intensity present in stained images from different sources [4].
Save the training and validation datastores to disk.
save(fullfile(dataDir,"trainingAndValidationDatastores.mat"),"dsTrainLabeled","dsValLabeled");
You can now use the datastores to train and validate a classification network. For an example, see Classify Tumors in Multiresolution Blocked Images.
References
[1] Ehteshami Bejnordi, Babak, Mitko Veta, Paul Johannes van Diest, Bram van Ginneken, Nico Karssemeijer, Geert Litjens, Jeroen A. W. M. van der Laak, et al. “Diagnostic Assessment of Deep Learning Algorithms for Detection of Lymph Node Metastases in Women With Breast Cancer.” JAMA 318, no. 22 (December 12, 2017): 2199–2210. https://doi.org/10.1001/jama.2017.14585.
[2] Szegedy, Christian, Vincent Vanhoucke, Sergey Ioffe, Jonathon Shlens, and Zbigniew Wojna. “Rethinking the Inception Architecture for Computer Vision,” December 2, 2015. https://arxiv.org/abs/1512.00567v3.
[3] ImageNet. https://www.image-net.org.
[4] Macenko, Marc, Marc Niethammer, J. S. Marron, David Borland, John T. Woosley, Xiaojun Guan, Charles Schmitt, and Nancy E. Thomas. “A Method for Normalizing Histology Slides for Quantitative Analysis.” In 2009 IEEE International Symposium on Biomedical Imaging: From Nano to Macro, 1107–10, 2009. https://doi.org/10.1109/ISBI.2009.5193250.
See Also
blockedImageDatastore | blockedImage | selectBlockLocations | mergeBlockLocationSets | blockLocationSet | bigimageshow | transform
Topics
- Classify Tumors in Multiresolution Blocked Images
- Set Up Spatial Referencing for Blocked Images
- Process Blocked Images Efficiently Using Partial Images or Lower Resolutions
- Process Blocked Images Efficiently Using Mask
- Create Labeled Blocked Image from ROIs and Masks
- Datastores for Deep Learning (Deep Learning Toolbox)