isoelectric
Estimate isoelectric point for amino acid sequence
Description
[
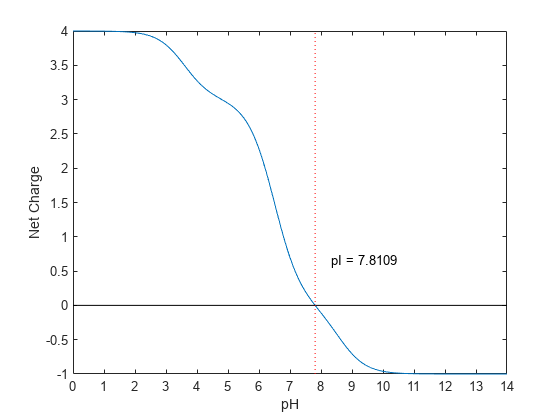
returns the estimated isoelectric point (pI,ChargePH] = isoelectric(SeqAA)pI) for an amino acid sequence
and the estimated charge for a given pH (default is typical intracellular pH
7.2).
The estimates are skewed by the underlying assumptions that all amino acids are fully
exposed to the solvent, that neighboring peptides have no influence on the pK of any given
amino acid, and that the constitutive amino acids, as well as the N- and C-termini, are
unmodified. Cysteine residues participating in disulfide bridges also affect the true
pI and are not considered here. By default,
isoelectric uses the EMBOSS amino acid pK table, or you can substitute
other values using the property PKVals.
If the sequence contains ambiguous amino acid characters (b z * –),
isoelectricignores the characters and displays a warning message.Warning: Symbols other than the standard 20 amino acids appear in the sequence.
If the sequence contains undefined amino acid characters (
i j o) ,isoelectricignores the characters and displays a warning message.Warning: Sequence contains unknown characters. These will be ignored.
___ = isoelectric(___,
specifies additional options using one or more name-value arguments. Use any arguments from
the previous syntaxes.Name=Value)
Examples
Input Arguments
Name-Value Arguments
Output Arguments
Version History
Introduced in R2006a