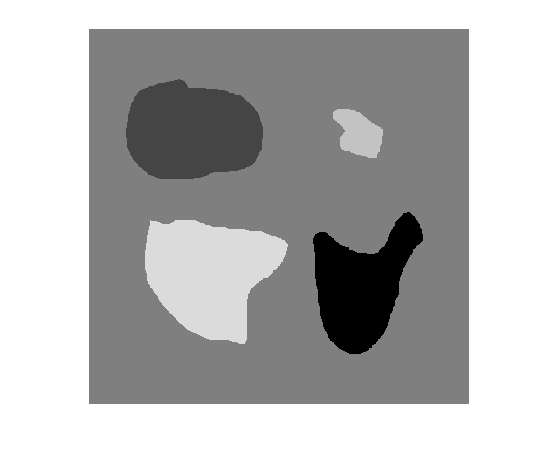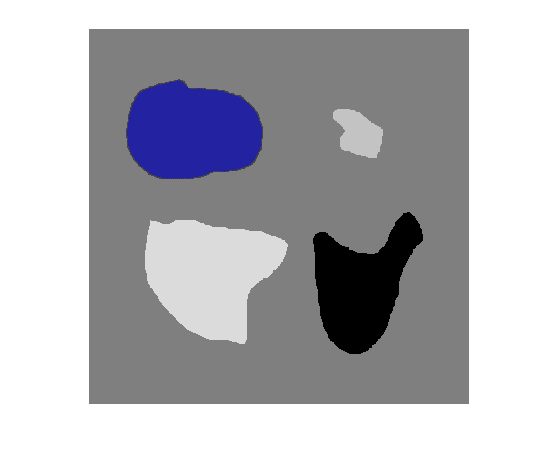activecontour
Segment image into foreground and background using active contours (snakes) region growing technique
Syntax
Description
The active contours technique, also called
snakes, is an iterative region-growing image segmentation
algorithm. Using the active contour algorithm, you specify initial curves on an image
and then use the activecontour function to evolve the curves
towards object boundaries.
BW = activecontour(A,mask)A into foreground (object) and background
regions using active contours.
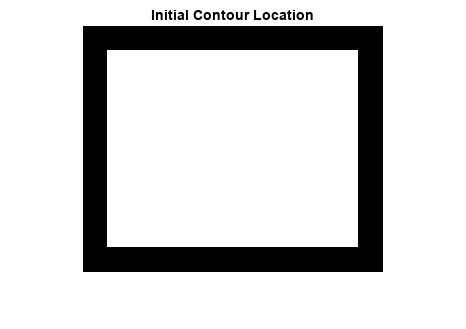
The mask argument is a binary image that specifies the
initial state of the active contour. The boundaries of the object regions (white) in
mask define the initial contour position used for contour
evolution to segment the image. The output image BW is a binary
image where the foreground is white (logical true) and the
background is black (logical false).
To obtain faster and more accurate segmentation results, specify an initial contour position that is close to the desired object boundaries.
BW = activecontour(___,Name=Value)
Examples
Input Arguments
Name-Value Arguments
Output Arguments
Tips
activecontouruses the boundaries of the regions inmaskas the initial state of the contour from where the evolution starts. Holes in the mask can cause unpredictable results. Useimfillto fill any holes in the regions inmask.If a region touches the image borders, then
activecontourremoves a single-pixel layer from the region, before further processing, so that the region does not touch the image border.To get faster and more accurate results, specify an initial contour position that is close to the desired object boundaries, especially for the
"edge"method.For the
"edge"method, the active contour is naturally biased towards shrinking inwards (collapsing). In the absence of any image gradient, the active contour shrinks on its own. Conversely, with the"Chan-Vese"method, where the contour is unbiased, the contour is free to either shrink or expand based on the image features.To achieve an accurate segmentation with the
"edge"method, specify an initial contour that lies outside the boundaries of the object. The active contour with the"edge"method is biased to shrink, by default.If object regions are of significantly different grayscale intensities, then the
"Chan-Vese"method [1] might not segment all objects in the image. For example, if the image contains objects that are brighter than the background and some that are darker, the"Chan-Vese"method typically segments out either the dark or the bright objects only.
Algorithms
activecontour uses the Sparse-Field level-set method, similar to the
method described in [3], for implementing active contour evolution.
References
[1] T. F. Chan, L. A. Vese, Active contours without edges. IEEE Transactions on Image Processing, Volume 10, Issue 2, pp. 266-277, 2001.
[2] V. Caselles, R. Kimmel, G. Sapiro, Geodesic active contours. International Journal of Computer Vision, Volume 22, Issue 1, pp. 61-79, 1997.
[3] R. T. Whitaker, A level-set approach to 3d reconstruction from range data. International Journal of Computer Vision, Volume 29, Issue 3, pp. 203-231, 1998.