simsmooth
State-space model simulation smoother
Description
X = simsmooth(Mdl,Y)X by applying a simulation
smoother to the time-invariant or time-varying state-space
model
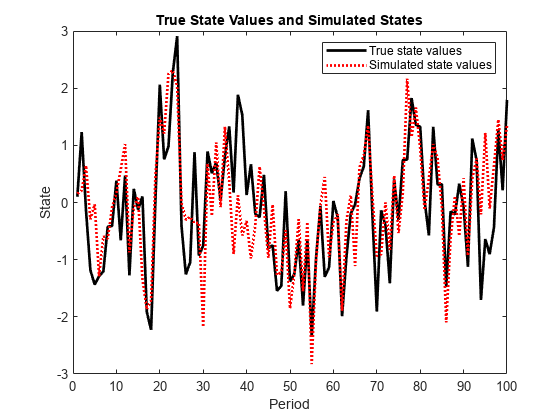
Mdl and responses Y. simsmooth
uses forward filtering and back sampling to obtain one random path from the posterior
distribution of the states.
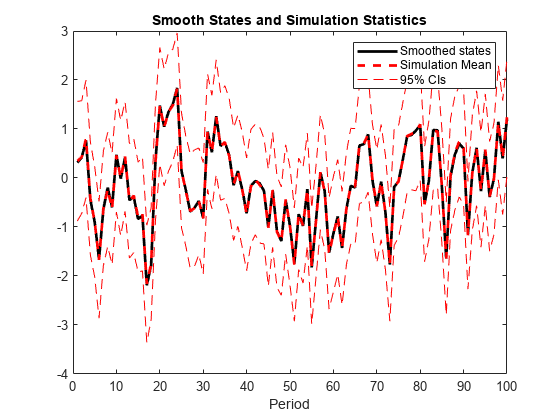
X = simsmooth(Mdl,Y,Name=Value)simsmooth(Mdl,Y,NumPaths=100) returns 100
independently generated paths of states.
Examples
Input Arguments
Name-Value Arguments
Output Arguments
More About
Algorithms
The Kalman filter accommodates missing data by not updating filtered state estimates corresponding to missing observations. In other words, suppose there is a missing observation at period t. Then, the state forecast for period t based on the previous t – 1 observations and filtered state for period t are equivalent.
For increased speed in simulating states, the simulation smoother implements minimal dimensionality error checking. Therefore, for models with unknown parameter values, you should ensure that the dimensions of the data and the dimensions of the coefficient matrices are consistent.
References
[1] Durbin J., and S. J. Koopman. “A Simple and Efficient Simulation Smoother for State Space Time Series Analysis.” Biometrika. Vol 89., No. 3, 2002, pp. 603–615.
Version History
Introduced in R2014b