infer
Infer univariate ARIMA or ARIMAX model residuals or conditional variances
Syntax
Description
Tbl2 = infer(Mdl,Tbl1)Tbl2 containing paths of
residuals and conditional variances inferred from the model
Mdl and the response data in the input table or
timetable Tbl1. (since R2023b)
infer selects the response variable named in
Mdl.SeriesName or the sole variable in
Tbl1. To select a different response variable in
Tbl1 to infer residuals and conditional variances, use
the ResponseVariable name-value argument.
[___] = infer(___,
specifies options using one or more name-value arguments in
addition to any of the input argument combinations in previous syntaxes.
Name=Value)infer returns the output argument combination for the
corresponding input arguments. For example, infer(Mdl,Y,Y0=PS,X=Pred) infers
residuals from the numeric vector of responses Y with
respect to the ARIMAX Mdl, and specifies the numeric vector
of presample response data PS to initialize the model and the
exogenous predictor data Pred for the regression
component.
Examples
Infer residuals from an AR model by supplying a hypothetical response series in a vector.
Specify an AR(2) model using known parameters.
Mdl = arima(AR={0.5 -0.8},Constant=0.002, ...
Variance=0.8);Simulate response data with 100 observations.
rng(1,"twister");
Y = simulate(Mdl,100);Y is a 100-by-1 vector containing a random response path drawn from Mdl.
Infer residuals for all corresponding responses.
E = infer(Mdl,Y);
E is a 100-by-1 vector containing a residuals corresponding to Y, with respect to Mdl. By default, infer backcasts for required presample observations.
Plot the residuals.
figure
plot(E)
title("Inferred Residuals")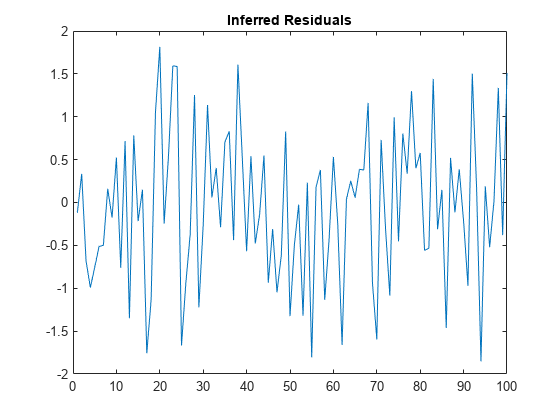
Infer the conditional variances from an AR(1) and GARCH(1,1) composite model. Return the loglikelihood value.
Specify an AR(1) model using known parameters. Set the variance equal to a garch model.
Mdl = arima(AR={0.8 -0.3},Constant=0);
MdlVar = garch(Constant=0.0002,GARCH=0.6,ARCH=0.2);
Mdl.Variance = MdlVar;Simulate response data with 100 observations.
rng(1,"twister")
Y = simulate(Mdl,100);Infer residuals and conditional variances for the entire response series. Compute the loglikelihood at the simulated data.
[E,V,logL] = infer(Mdl,Y); logL
logL = 209.6405
E and V are 100-by-1 vectors of inferred residuals and conditional variances, given the response data and model.
Plot the conditional variances.
figure
plot(V)
title("Inferred Conditional Variances")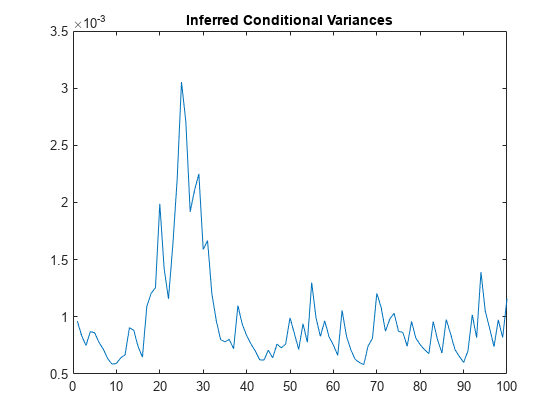
Infer residuals from an AR model by supplying a hypothetical response series in a vector. Supply presample responses to initialize the model.
Specify an AR(2) model using known parameters.
Mdl = arima(AR={0.5 -0.8},Constant=0.002, ...
Variance=0.8)Mdl =
arima with properties:
Description: "ARIMA(2,0,0) Model (Gaussian Distribution)"
SeriesName: "Y"
Distribution: Name = "Gaussian"
P: 2
D: 0
Q: 0
Constant: 0.002
AR: {0.5 -0.8} at lags [1 2]
SAR: {}
MA: {}
SMA: {}
Seasonality: 0
Beta: [1×0]
Variance: 0.8
Consider inferring residuals from a response series of length T = 100. Because the model requires Mdl.P responses to initialize the model, simulate T + Mdl.P = 102 responses from the model.
rng(1,"twister");
T = 100;
TSim = T + Mdl.P;
y = simulate(Mdl,TSim);Y is a 102-by-1 vector representing a random response path drawn from the model.
Infer residuals from the last T response and use the first Mdl.P observations as a presample to initialize the model.
E = infer(Mdl,y((Mdl.P+1):end),Y0=y(1:Mdl.P)); size(E)
ans = 1×2
100 1
E is a 100-by-1 vector containing a residuals corresponding to the last 100 observations of y, with respect to Mdl.
Plot the residuals.
figure
plot(E)
title("Inferred Residuals")
Since R2023b
Fit an ARIMA(1,1,1) model to the weekly average NYSE closing prices. Supply timetables of in-sample and presample data for the fit. Then, infer the residuals from the fit.
Load Data
Load the US equity index data set Data_EquityIdx.
load Data_EquityIdx
T = height(DataTimeTable)T = 3028
The timetable DataTimeTable includes the time series variable NYSE, which contains daily NYSE composite closing prices from January 1990 through December 2001.
Plot the daily NYSE price series.
figure
plot(DataTimeTable.Time,DataTimeTable.NYSE)
title("NYSE Daily Closing Prices: 1990 - 2001")
Prepare Timetable for Estimation
When you plan to supply a timetable, you must ensure it has all the following characteristics:
The selected response variable is numeric and does not contain any missing values.
The timestamps in the
Timevariable are regular, and they are ascending or descending.
Remove all missing values from the timetable, relative to the NYSE price series.
DTT = rmmissing(DataTimeTable,DataVariables="NYSE");
T_DTT = height(DTT)T_DTT = 3028
Because all sample times have observed NYSE prices, rmmissing does not remove any observations.
Determine whether the sampling timestamps have a regular frequency and are sorted.
areTimestampsRegular = isregular(DTT,"days")areTimestampsRegular = logical
0
areTimestampsSorted = issorted(DTT.Time)
areTimestampsSorted = logical
1
areTimestampsRegular = 0 indicates that the timestamps of DTT are irregular. areTimestampsSorted = 1 indicates that the timestamps are sorted. Business day rules make daily macroeconomic measurements irregular.
Remedy the time irregularity by computing the weekly average closing price series of all timetable variables.
DTTW = convert2weekly(DTT,Aggregation="mean"); areTimestampsRegular = isregular(DTTW,"weeks")
areTimestampsRegular = logical
1
T_DTTW = height(DTTW)
T_DTTW = 627
DTTW is regular.
figure
plot(DTTW.Time,DTTW.NYSE)
title("NYSE Daily Closing Prices: 1990 - 2001")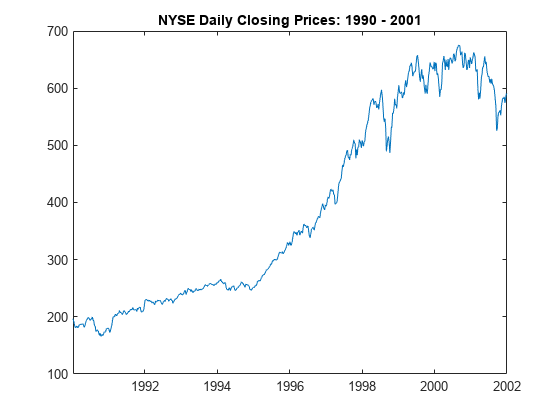
Create Model Template for Estimation
Suppose that an ARIMA(1,1,1) model is appropriate to model NYSE composite series during the sample period.
Create an ARIMA(1,1,1) model template for estimation.
Mdl = arima(1,1,1);
Mdl is a partially specified arima model object.
Fit Model to Data
infer requires Mdl.P presample observations to initialize the model. infer backcasts for necessary presample responses, but you can provide a presample.
Partition the data into presample and in-sample, or estimation sample, observations.
T0 = Mdl.P; DTTW0 = DTTW(1:T0,:); DTTW1 = DTTW((T0+1):end,:);
Fit an ARIMA(1,1,1) model to the in-sample weekly average NYSE closing prices. Specify the response variable name, presample timetable, and the presample response variable name.
EstMdl = estimate(Mdl,DTTW1,ResponseVariable="NYSE", ... Presample=DTTW0,PresampleResponseVariable="NYSE");
ARIMA(1,1,1) Model (Gaussian Distribution):
Value StandardError TStatistic PValue
________ _____________ __________ ___________
Constant 0.83623 0.453 1.846 0.064892
AR{1} -0.32862 0.23526 -1.3968 0.16247
MA{1} 0.42703 0.22613 1.8884 0.058967
Variance 56.065 1.8433 30.416 3.3798e-203
EstMdl is a fully specified, estimated arima model object.
Infer Residuals
Infer the residuals from the fitted model and in-sample observations. Specify the response variable name, presample timetable, and the presample response variable name.
Tbl2 = infer(EstMdl,DTTW1,ResponseVariable="NYSE", ... Presample=DTTW0,PresampleResponseVariable="NYSE"); tail(Tbl2)
Time NYSE NASDAQ Y_Residual Y_Variance
___________ ______ ______ __________ __________
16-Nov-2001 577.11 1886.9 5.8649 56.065
23-Nov-2001 583 1898.3 5.3303 56.065
30-Nov-2001 581.41 1925.8 -2.7678 56.065
07-Dec-2001 584.96 1998.1 3.3787 56.065
14-Dec-2001 574.03 1981 -12.038 56.065
21-Dec-2001 582.1 1967.9 8.7774 56.065
28-Dec-2001 590.28 1967.2 6.2526 56.065
04-Jan-2002 589.8 1950.4 -1.3008 56.065
size(Tbl2)
ans = 1×2
625 4
Tbl2 is a 625-by-4 timetable containing all variables in DTTW1, and the inferred residuals from the fit NYSE_Response and constant variance paths NYSE_Variance (Mdl.Variance = 56.065).
Since R2023b
Fit an ARIMA(1,1,1) model to the weekly average NYSE closing prices. Supply a timetable of data and specify the series for the fit. Then, compute fitted responses.
Load the US equity index data set Data_EquityIdx.
load Data_EquityIdx
T = height(DataTimeTable)T = 3028
Remedy the time irregularity by computing the weekly average closing price series of all timetable variables.
DTTW = convert2weekly(DataTimeTable,Aggregation="mean");
T_DTTW = height(DTTW)T_DTTW = 627
Create an ARIMA(1,1,1) model template for estimation. Set the response series name to NYSE.
Mdl = arima(1,1,1);
Mdl.SeriesName = "NYSE";Partition the data into presample and in-sample, or estimation sample, observations.
T0 = Mdl.P; DTTW0 = DTTW(1:T0,:); DTTW1 = DTTW((T0+1):end,:);
Fit an ARIMA(1,1,1) model to the in-sample weekly average NYSE closing prices. Specify the presample timetable, and the presample response variable name.
EstMdl = estimate(Mdl,DTTW1,Presample=DTTW0, ... PresampleResponseVariable="NYSE");
ARIMA(1,1,1) Model (Gaussian Distribution):
Value StandardError TStatistic PValue
________ _____________ __________ ___________
Constant 0.83623 0.453 1.846 0.064891
AR{1} -0.32862 0.23526 -1.3968 0.16247
MA{1} 0.42703 0.22613 1.8884 0.058966
Variance 56.065 1.8433 30.416 3.3799e-203
Infer the residuals from the fitted model and in-sample observations. Specify the presample timetable, and the presample response variable name.
Tbl2 = infer(EstMdl,DTTW1,Presample=DTTW0, ... PresampleResponseVariable="NYSE");
Compute fitted response values by subtracting the residuals from the observed response series.
Tbl2.YHat = Tbl2.NYSE - Tbl2.NYSE_Residual;
Plot the observed responses and the fitted values.
figure plot(Tbl2.Time,[Tbl2.NYSE Tbl2.YHat]) legend("Observations","Fitted values") title("NYSE Weekly Average Price Series")
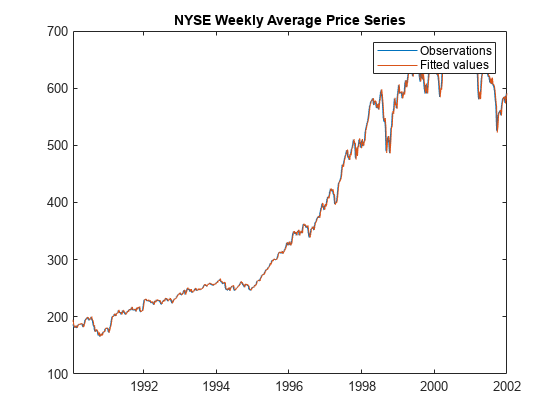
The fitted values closely track the observations.
Plot the residuals versus the fitted values.
figure plot(Tbl2.YHat,Tbl2.NYSE_Residual,".",MarkerSize=15) ylabel("Residuals") xlabel("Fitted Values") title("Residual Plot")
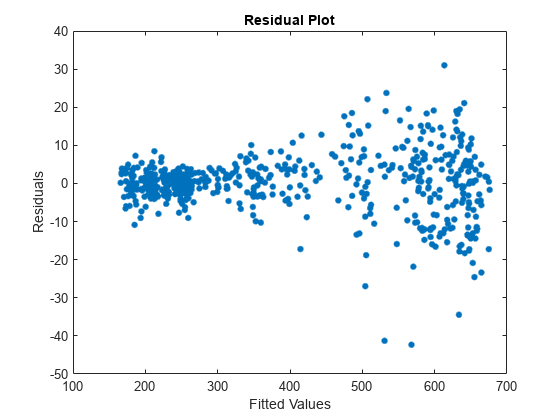
Residual variance appears larger for larger fitted values. One remedy for this behavior is to apply the log transform to the data.
Infer residuals from an ARMAX model.
Specify an ARMA(1,2) model using known parameters for the response (MdlY) and an AR(1) model for the predictor data (MdlX).
MdlY = arima(AR=0.2,MA={-0.1,0.6},Constant=1, ...
Variance=2,Beta=3)MdlY =
arima with properties:
Description: "ARIMAX(1,0,2) Model (Gaussian Distribution)"
SeriesName: "Y"
Distribution: Name = "Gaussian"
P: 1
D: 0
Q: 2
Constant: 1
AR: {0.2} at lag [1]
SAR: {}
MA: {-0.1 0.6} at lags [1 2]
SMA: {}
Seasonality: 0
Beta: [3]
Variance: 2
MdlX = arima(AR=0.3,Constant=0,Variance=1);
If you do not specify presample responses, infer requires at least T + MdlY.P predictor observations to simulate a response series of length T.
Consider simulating a response series of length 100. Simulate a predictor series of length 101, and then simulate the response series. Provide the predictor data to simulate for the exogenous regression component.
rng(1,"twister") % For reproducibility T = 100; Pred = simulate(MdlX,T + MdlY.P); Y = simulate(MdlY,T,X=Pred);
Infer residuals using the entire series.
E = infer(MdlY,Y,X=Pred);
figure
plot(E)
title("Inferred Residuals") 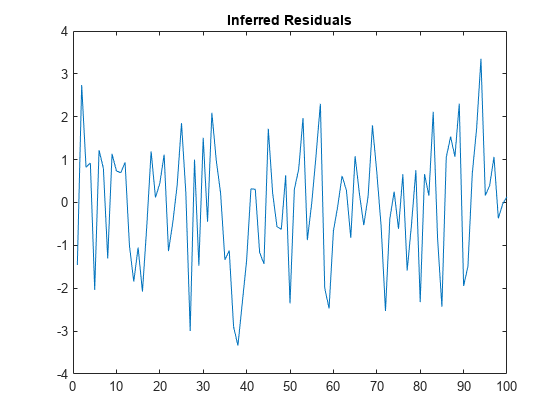
Input Arguments
Response data yt, specified as a
numobs-by-1 numeric column vector or
numobs-by-numpaths numeric matrix.
numObs is the length of the time series (sample
size). numpaths is the number of separate, independent
paths of response series.
infer infers the residuals and conditional
variances of columns of Y, which are time series
characterized by Mdl. Y is the
continuation of the presample series Y0.
Each row corresponds to a sampling time. The last row contains the latest set of observations.
Each column corresponds to a separate, independent path of response data.
infer assumes that responses across any row
occur simultaneously.
Data Types: double
Since R2023b
Time series data containing the observed response variable
yt and, optionally,
predictor variables xt for the
exogenous regression component, specified as a table or timetable with
numvars variables and numobs rows.
You can optionally select the response variable or
numpreds predictor variables by using the
ResponseVariable or
PredictorVariables name-value arguments,
respectively.
Each row is an observation, and measurements in each row occur
simultaneously. The selected response variable is a single path
(numobs-by-1 vector) or multiple paths
(numobs-by-numpaths matrix) of
numobs observations of response data.
Each path (column) of the selected response variable is independent of the
other paths, but path jjnumpaths. Each selected predictor variable is a
numobs-by-1 numeric vector representing one path. The
infer function includes all predictor
variables in the model when it infers residuals and conditional variances.
Variables in Tbl1 represent the continuation of
corresponding variables in Presample.
If Tbl1 is a timetable, it must represent a sample
with a regular datetime time step (see isregular), and the datetime
vector Tbl1.Time must be strictly ascending or
descending.
If Tbl1 is a table, the last row contains the latest
observation.
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: infer(Mdl,Y,Y0=PS,X=Pred) infers residuals from the
numeric vector of responses Y through the ARIMAX
Mdl, and specifies the numeric vector of presample response
data PS to initialize the model and the exogenous predictor data
Pred for the regression component.
Since R2023b
Response variable yt to select from
Tbl1 containing the response data, specified as one of the
following data types:
String scalar or character vector containing a variable name in
Tbl1.Properties.VariableNamesVariable index (positive integer) to select from
Tbl1.Properties.VariableNamesA logical vector, where
DisturbanceVariable(selects variablej) = truejTbl1.Properties.VariableNames
The selected variable must be a numeric vector and cannot contain missing values
(NaNs).
If Tbl1 has one variable, the default specifies that variable.
Otherwise, the default matches the variable to names in
Mdl.SeriesName.
Example: ResponseVariable="StockRate"
Example: ResponseVariable=[false false true false] or
ResponseVariable=3 selects the third table variable as the
response variable.
Data Types: double | logical | char | cell | string
Presample response data yt
to initialize the model, specified as a
numpreobs-by-1 numeric column vector or a
numpreobs-by-numprepaths
numeric matrix. Use Y0 only when you supply the
numeric array of response data Y.
numpreobs is the number of presample observations.
numprepaths is the number of presample response
paths.
Each row is a presample observation (sampling time), and measurements
in each row occur simultaneously. The last row contains the latest
presample observation. numpreobs must be at least
Mdl.P to initialize the AR model component. If
numpreobs > Mdl.P,
infer uses the latest required number of
observations only.
Columns of Y0 are separate, independent presample
paths. The following conditions apply:
If
Y0is a column vector, it represents a single response path.inferapplies it to each output path.If
Y0is a matrix, each column represents a presample response path.inferappliesY0(:,to initialize pathj)j.numprepathsmust be at leastnumpaths. Ifnumprepaths>numpaths,inferuses the firstsize(Y,2)columns only.
By default, infer backcasts to obtain the
necessary observations.
Data Types: double
Presample residual data et
to initialize the model, specified as a
numpreobs-by-1 numeric column vector or a
numpreobs-by-numprepaths
numeric matrix. Use E0 only when you supply the
numeric array of response data Y.
Each row is a presample observation (sampling time), and measurements
in each row occur simultaneously. The last row contains the latest
presample observation. numpreobs must be at least
Mdl.Q to initialize the MA model component. If
Mdl.Variance is a conditional variance model (for
example, a garch model object),
infer can require more rows than
Mdl.Q. If numpreobs is larger
than required, infer uses the latest required
number of observations only.
Columns of E0 are separate, independent presample
paths. The following conditions apply:
If
E0is a column vector, it represents a single residual path.inferapplies it to each output path.If
E0is a matrix, each column represents a presample residual path.inferappliesE0(:,to initialize pathj)j.numprepathsmust be at leastnumpaths. Ifnumprepaths>numpaths,inferuses the firstsize(Y,2)columns only.inferassumes each column ofE0has a mean of zero.
By default, infer sets the necessary
presample disturbances to zero.
Data Types: double
Presample conditional variances
σt2
to initialize the conditional variance model, specified as a
numpreobs-by-1 positive numeric column vector or
a numpreobs-by-numprepaths
positive numeric matrix. If the conditional variance
Mdl.Variance is constant,
infer ignores V0. Use
V0 only when you supply the numeric array of
response data Y.
Each row is a presample observation (sampling time), and measurements
in each row occur simultaneously. The last row contains the latest
presample observation. numpreobs must be at least
Mdl.Q to initialize the conditional variance
model in Mdl.Variance. For details, see the infer function of
conditional variance models. If numpreobs is larger
than required, infer uses the latest required
number of observations only.
Columns of V0 are separate, independent presample
paths. The following conditions apply:
If
V0is a column vector, it represents a single path of conditional variances.inferapplies it to each output path.If
V0is a matrix, each column represents a presample path of conditional variances.inferappliesV0(:,to initialize pathj)j.numprepathsmust be at leastnumpaths. Ifnumprepaths>numpaths,inferuses the firstsize(Y,2)columns only.
By default, infer sets all necessary
presample conditional variances to the average squared value of inferred
residuals.
Data Types: double
Since R2023b
Presample data containing paths of response
yt, residual
et, or conditional
variance
σt2
series to initialize the model, specified as a table or timetable, the
same type as Tbl1, with
numprevars variables and
numpreobs rows. Use
Presample only when you supply a table or
timetable of data Tbl1.
Each selected variable is a single path
(numpreobs-by-1 vector) or multiple paths
(numpreobs-by-numprepaths
matrix) of numpreobs observations representing the
presample of the response, residual, or conditional variance series for
ResponseVariable, the selected response
variable in Tbl1.
Each row is a presample observation, and measurements in each row
occur simultaneously. numpreobs must be one of the
following values:
At least
Mdl.PwhenPresampleprovides only presample responsesAt least
Mdl.QwhenPresampleprovides only presample disturbances or conditional variancesAt least
max([Mdl.P Mdl.Q])otherwise
When Mdl.Variance is a conditional
variance model, infer can require more than
the minimum required number of presample values.
If you supply more rows than necessary,
infer uses the latest required number of
observations only.
When Presample provides presample residuals,
infer assumes each presample residual
path has a mean of zero.
If Presample is a timetable, all the following
conditions must be true:
Presamplemust represent a sample with a regular datetime time step (seeisregular).The inputs
Tbl1andPresamplemust be consistent in time such thatPresampleimmediately precedesTbl1with respect to the sampling frequency and order.The datetime vector of sample timestamps
Presample.Timemust be ascending or descending.
If Presample is a table, the last row contains
the latest presample observation.
By default:
When
Mdlis a model without a exogenous linear regression component (ARIMAX),inferbackcasts for necessary presample responses, sets necessary presample residuals to 0, and sets necessary presample variances to the average squared value of inferred residuals.When
Mdlis an ARIMAX model (you specify thePredictorVariablesname-value argument), you must specify presample response data, butinfersets necessary presample residuals to 0 and sets necessary presample variances to the average squared value of inferred residuals.
If you specify the Presample, you must specify
the presample response, residual, or conditional variance name by using
the PresampleResponseVariable,
PresampleInnovationVariable, or
PresampleVarianceVariable name-value
argument.
Since R2023b
Response variable yt to select from
Presample containing presample response data, specified as one of
the following data types:
String scalar or character vector containing a variable name in
Presample.Properties.VariableNamesVariable index (positive integer) to select from
Presample.Properties.VariableNamesA logical vector, where
PresampleResponseVariable(selects variablej) = truejPresample.Properties.VariableNames
The selected variable must be a numeric matrix and cannot contain missing values
(NaNs).
If you specify presample response data by using the Presample
name-value argument, you must specify
PresampleResponseVariable.
Example: PresampleResponseVariable="Stock0"
Example: PresampleResponseVariable=[false false true false] or
PresampleResponseVariable=3 selects the third table variable as
the presample response variable.
Data Types: double | logical | char | cell | string
Since R2023b
Presample residual variable
et to select from
Presample containing presample residual data,
specified as one of the following data types:
String scalar or character vector containing a variable name in
Presample.Properties.VariableNamesVariable index (positive integer) to select from
Presample.Properties.VariableNamesA logical vector, where
PresampleInnovationVariable(selects variablej) = truejPresample.Properties.VariableNames
The selected variable must be a numeric matrix and cannot contain
missing values (NaNs).
If you specify presample residual data by using the
Presample name-value argument, you must specify
PresampleInnovationVariable.
Example: PresampleInnovationVariable="StockRateDist0"
Example: PresampleInnovationVariable=[false false true
false] or PresampleInnovationVariable=3
selects the third table variable as the presample innovation
variable.
Data Types: double | logical | char | cell | string
Since R2023b
Conditional variance variable
σt2 to select
from Presample containing presample conditional variance data,
specified as one of the following data types:
String scalar or character vector containing a variable name in
Presample.Properties.VariableNamesVariable index (positive integer) to select from
Presample.Properties.VariableNamesA logical vector, where
PresampleVarianceVariable(selects variablej) = truejPresample.Properties.VariableNames
The selected variable must be a numeric vector and cannot contain missing values
(NaNs).
If you specify presample conditional variance data by using the
Presample name-value argument, you must specify
PresampleVarianceVariable.
Example: PresampleVarianceVariable="StockRateVar0"
Example: PresampleVarianceVariable=[false false true false] or
PresampleVarianceVariable=3 selects the third table variable as
the presample conditional variance variable.
Data Types: double | logical | char | cell | string
Exogenous predictor data for the model regression component, specified
as a numeric matrix with numpreds columns.
numpreds is the number of predictor variables
(numel(Mdl.Beta)). Use X only
when you supply the numeric array of response data
Y.
If you do not specify Y0, the number of rows of
X must be at least numObs +
Mdl.P. Otherwise, the number of rows of
X must be at least numObs. If
the number of rows of X exceeds the number necessary,
infer uses only the latest observations.
infer does not use the regression
component in the presample period.
Columns of X are separate predictor
variables.
infer applies X to each
path; that is, X represents one path of observed
predictors.
By default, infer excludes the regression
component, regardless of its presence in
Mdl.
Data Types: double
Since R2023b
Exogenous predictor variables
xt to select from
Tbl1 containing the predictor data for the model
regression component, specified as one of the following data types:
String vector or cell vector of character vectors containing
numpredsvariable names inTbl1.Properties.VariableNamesA vector of unique indices (positive integers) of variables to select from
Tbl1.Properties.VariableNamesA logical vector, where
PredictorVariables(selects variablej) = truejTbl1.Properties.VariableNames
The selected variables must be numeric vectors and cannot contain
missing values (NaNs).
If you specify PredictorVariables, you must also
specify presample response data to by using the
Presample and
PresampleResponseVariable name-value arguments.
For more details, see Algorithms.
By default, infer excludes the regression
component, regardless of its presence in
Mdl.
Example: PredictorVariables=["M1SL" "TB3MS"
"UNRATE"]
Example: PredictorVariables=[true false true false]
or PredictorVariable=[1 3] selects the first and
third table variables to supply the predictor data.
Data Types: double | logical | char | cell | string
Note
NaNvalues inY,X,Y0,E0, andV0indicate missing values.inferremoves missing values from specified data by list-wise deletion.For the presample,
inferhorizontally concatenates the possibly jagged arraysY0,E0, andV0with respect to the last rows, and then it removes any row of the concatenated matrix containing at least oneNaN.For in-sample data,
inferhorizontally concatenates the possibly jagged arraysYandX, and then it removes any row of the concatenated matrix containing at least oneNaN.
This type of data reduction reduces the effective sample size and can create an irregular time series.
For numeric data inputs,
inferassumes that you synchronize the presample data such that the latest observations occur simultaneously.inferissues an error when any table or timetable input contains missing values.
Output Arguments
Inferred residual paths et,
returned as a numobs-by-numpaths
numeric matrix. infer returns
E only when you supply the input
Y.
E(
is the path j,k)kj; it is the residual associated with
response
Y(.j,k)
Inferred conditional variance paths
σt, returned as a
numobs-by-numpaths numeric matrix.
infer returns V only
when you supply the input Y.
V(
is the path j,k)kj; it is the conditional
variance associated with response
Y(.j,k)
Since R2023b
Inferred residual et and
conditional variance
σt2
paths, returned as a table or timetable, the same data type as
Tbl1. infer returns
Tbl2 only when you supply the input
Tbl1.
Tbl2 contains the following variables:
The inferred residual paths, which are in a
numobs-by-numpathsnumeric matrix, with rows representing observations and columns representing independent paths. Each path corresponds to the input response path inTbl1and represents the continuation of the corresponding presample residual path inPresample.infernames the inferred residual variable inTbl2responseName_ResidualresponseNameMdl.SeriesName. For example, ifMdl.SeriesNameisStockReturns,Tbl2contains a variable for the corresponding inferred innovations paths with the nameStockReturns_Residual.The inferred conditional variance paths, which are in a
numobs-by-numpathsnumeric matrix, with rows representing observations and columns representing independent paths. Each path represents the continuation of the corresponding path of presample conditional variances inPresample.infernames the inferred conditional variance variable inTbl2responseName_VarianceresponseNameMdl.SeriesName. For example, ifMdl.SeriesNameisStockReturns,Tbl2contains a variable for the corresponding inferred conditional variance paths with the nameStockReturns_Variance.All variables
Tbl1.
If Tbl1 is a timetable, row times of
Tbl1 and Tbl2 are
equal.
Algorithms
If you supply data in the table or timetable Tbl1 to estimate an
ARIMAX model, infer cannot backcast for presample responses.
Therefore, if you specify PredictorVariables, you must also specify
presample response data by using the Presample and
PresampleResponseVariable name-value arguments.
References
[1] Box, G. E. P., G. M. Jenkins, and G. C. Reinsel. Time Series Analysis: Forecasting and Control 3rd ed. Englewood Cliffs, NJ: Prentice Hall, 1994.
[2] Enders, W. Applied Econometric Time Series. Hoboken, NJ: John Wiley & Sons, 1995.
[3] Hamilton, J. D. Time Series Analysis. Princeton, NJ: Princeton University Press, 1994.
Version History
Introduced in R2012aIn addition to accepting input data (in-sample and presample) in numeric arrays,
infer accepts input data in tables or regular
timetables. When you supply data in a table or timetable, the following conditions
apply:
inferchooses the default in-sample response series on which to operate, but you can use the specified optional name-value argument to select a different series.If you specify optional presample response, residual, or conditional variance data to initialize the model, you must also specify the appropriate presample variable names.
inferreturns results in a table or timetable.
Name-value arguments to support tabular workflows include:
ResponseVariablespecifies the variable name of the response paths in the input data, from whichinferinfers conditional variances and innovations.Presamplespecifies the input table or timetable of presample innovation and conditional variance data.PresampleResponseVariablespecifies the variable name of the response paths to select fromPresample.PresampleInnovationVariablespecifies the variable name of the residual paths to select fromPresample.PresampleVarianceVariablespecifies the variable name of the conditional variance paths to select fromPresample.PredictorVariablesspecifies the names of the predictor series to select from the input data for a model regression component.
See Also
Objects
Functions
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)