mhsample
Generate Markov chain sample using Metropolis–Hastings sampler
Description
sample = mhsample(start,numsamples,Name=Value)numsamples random draws from the
target stationary distribution by using the Metropolis–Hastings sampler, a Markov chain
Monte Carlo (MCMC) algorithm. mhsample initializes the sampler from
start, and then draws samples using the proposal random number
generator.
You must specify these inputs using name-value arguments:
Target PDF Form — Specify the target stationary distribution or its log by using
PDForLogPDF, respectively.Proposal Random Number Generator — Specify a supported distribution name or function by using
ProposalorPropRND, respectively.Proposal Distribution Description — When you use
Proposal, specify the proposal scale usingScale. When you usePropRND, specify one of the following:Symmetricto indicate whether the proposal distribution is symmetricPropPDFto specify the proposal probability distribution function (pdf)LogPropPDFto specify the log of the proposal pdf
You can specify additional options by setting more name-value arguments. For example,
BurnIn=100,Thin=4 removes the first 100 draws from the raw generated
Markov chain (burn-in period of 100), retains every fourth draw of the remaining chain
(thinning factor of 4), and then returns the resulting processed chain in
sample.
For details on which Metropolis–Hastings sampler algorithm
mhsample implements, see Algorithms.
[
also returns the acceptance rate sample,acceptance] = mhsample(___)acceptance of proposal draws for each
Markov chain. When computing acceptance,
mhsample includes any samples omitted for the specified thinning
factor and burn-in period.
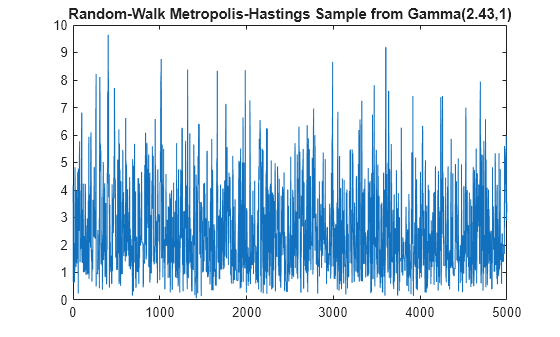
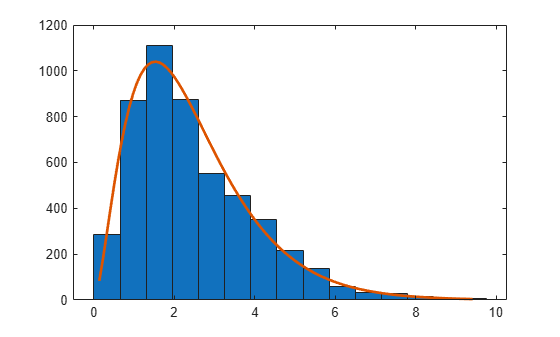
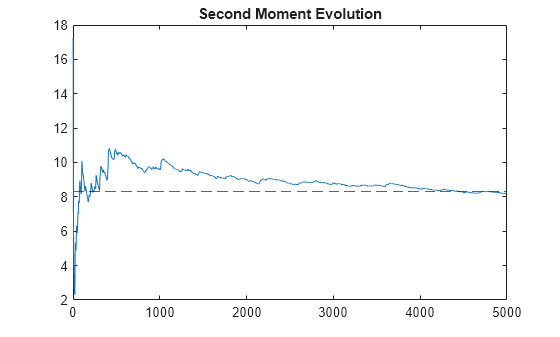
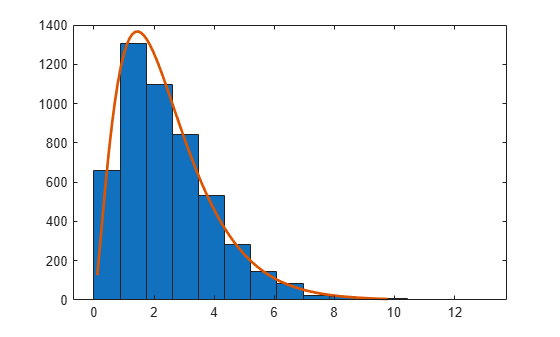
Examples
Input Arguments
Name-Value Arguments
Output Arguments
More About
Tips
When you plan to return multiple independent samples by using the
NumChainsname-value argument, ensure that all custom functions you pass tomhsamplereturn a numeric column vector or matrix of values, with each row corresponding to the values required by the corresponding Markov chain. The noncustom proposal distributions, available with theProposalname-value argument, enable you to avoid writing custom functions which might not return values in the correct form for multiple Markov chains. The noncustom proposal distributions are easy to use, provide increased sampling speed compared to custom proposals, can be tuned easily, and work well for a variety of applications. Therefore, a good practice is to try settingProposalbefore writing and setting custom proposal distributions.An acceptance rate close to 0 indicates that most proposal draws likely correspond to regions of low probability in the target distribution. An acceptance rate close to 1 can indicate that the proposal draws do not sufficiently explore the target distribution. However, an ideal acceptance rate depends on the target distribution. In general, Gelman, et al, suggest tuning the sampler until the acceptance rate is close to 0.25[1].
Algorithms
mhsample implements the Random-Walk or Independent
Metropolis–Hastings sampler algorithm depending on your specifications.
| Algorithm | Specifications |
|---|---|
| Random-Walk Metropolis–Hastings |
|
| Independent Metropolis–Hastings |
When using the custom proposal functions |
For each of NumChains generated Markov chains,
mhsample draws BurnIn +
Thin*numsamples samples, and
then processes each sample by removing the burn-in draws specified by
BurnIn, and then reducing the remaining draws as specified by the
thinning factor Thin.
This figure illustrates how mhsample reduces the raw MCMC sample.
Rectangles represent successive draws from the distribution. mhsample
removes the white rectangles from the Monte Carlo sample. The remaining
numsamples black rectangles compose sample.

References
[1] Gelman, Andrew, Walter R. Gilks, and Gareth O. Roberts. "Weak Convergence and Optimal Scaling of Random Walk Metropolis Algorithms." The Annals of Applied Probability. 7 (February 1997): 110–120. https://doi.org/10.1214/aoap/1034625254.