Explore 3-D Volumetric Data with Volume Viewer App
This example shows how to look at and explore 3-D volumetric data using the Volume Viewer app. Volume rendering is highly dependent on defining an appropriate alphamap so that structures in your data that you want to see are opaque and structures that you do not want to see are transparent. To illustrate, the example loads an MRI study of the human head into the Volume Viewer app and explores the data using the visualization capabilities of the Volume Viewer.
Load Volume Data into Volume Viewer
This part of the example shows how to load volumetric data into the Volume Viewer app.
Load the MRI data of a human head from a MAT-file into the workspace. The MRI data is a modified subset of the BraTS data set [1]. This operation creates a variable named D in your workspace that contains the volumetric data. Use the squeeze command to remove the singleton dimension from the data.
load mri
D = squeeze(D);
whosName Size Bytes Class Attributes D 128x128x27 442368 uint8 map 89x3 2136 double siz 1x3 24 double
Open the Volume Viewer app. From the MATLAB® toolstrip, open the Apps tab and under Image Processing and Computer Vision, click  . You can also open the app using the
. You can also open the app using the volumeViewer command.
volumeViewer(D)
Load volumetric data into the Volume Viewer app. Click Import Volume. You can load an image by specifying its file name or load a variable from the workspace. If you have volumetric data in a DICOM format that uses multiple files to represent a volume, you can specify the DICOM folder name. Choose the Import From Workspace option because the data is in the workspace.
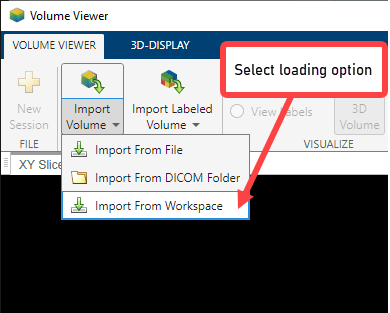
Select the workspace variable in the Import Volume dialog box and click OK.

To start a new instance of the Volume Viewer app, click New Session.
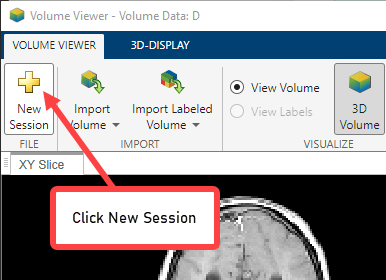
When you create a new session, this option deletes all the data currently in the viewer. Click Yes to create the new session.
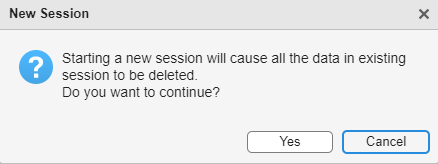
View Volume Data in Volume Viewer
In this part of the example, you decide how you want to view your data. Volume Viewer offers several options.
View the volume in the Volume Viewer app. Volume Viewer displays the data as a volume and as slice planes. The MRI data displayed as a volume is recognizable as a human head. To explore the volume, zoom in and out on the image using the mouse wheel or a right-click. You can also rotate the volume by positioning the cursor in the image window, pressing and holding the mouse, and moving the cursor. You are always zooming or rotating around the center of the volume. The position of the axes in the Orientation Axes window reflects the spatial orientation of the image as you rotate it.

To change the background color used in the display window, open the 3D-Display tab, click Background Color, and select a color.
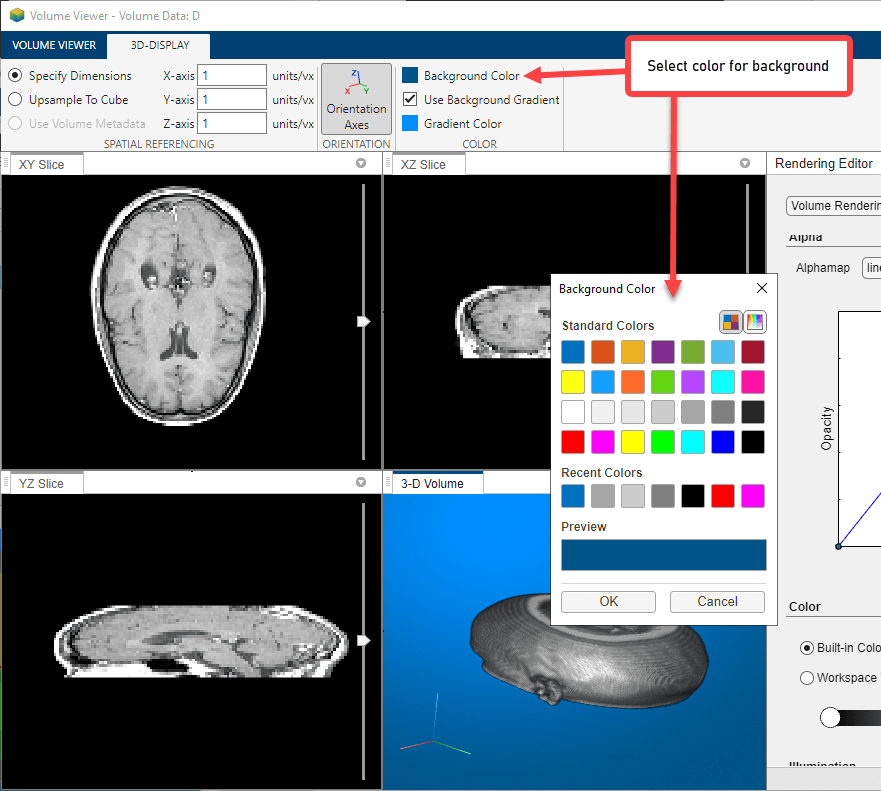
View the MRI data as a set of slice planes. Click Slice Planes. You can also zoom in and rotate this view of the data. Use the scroll functionality in the three slice windows to view individual slices in any of the planes.
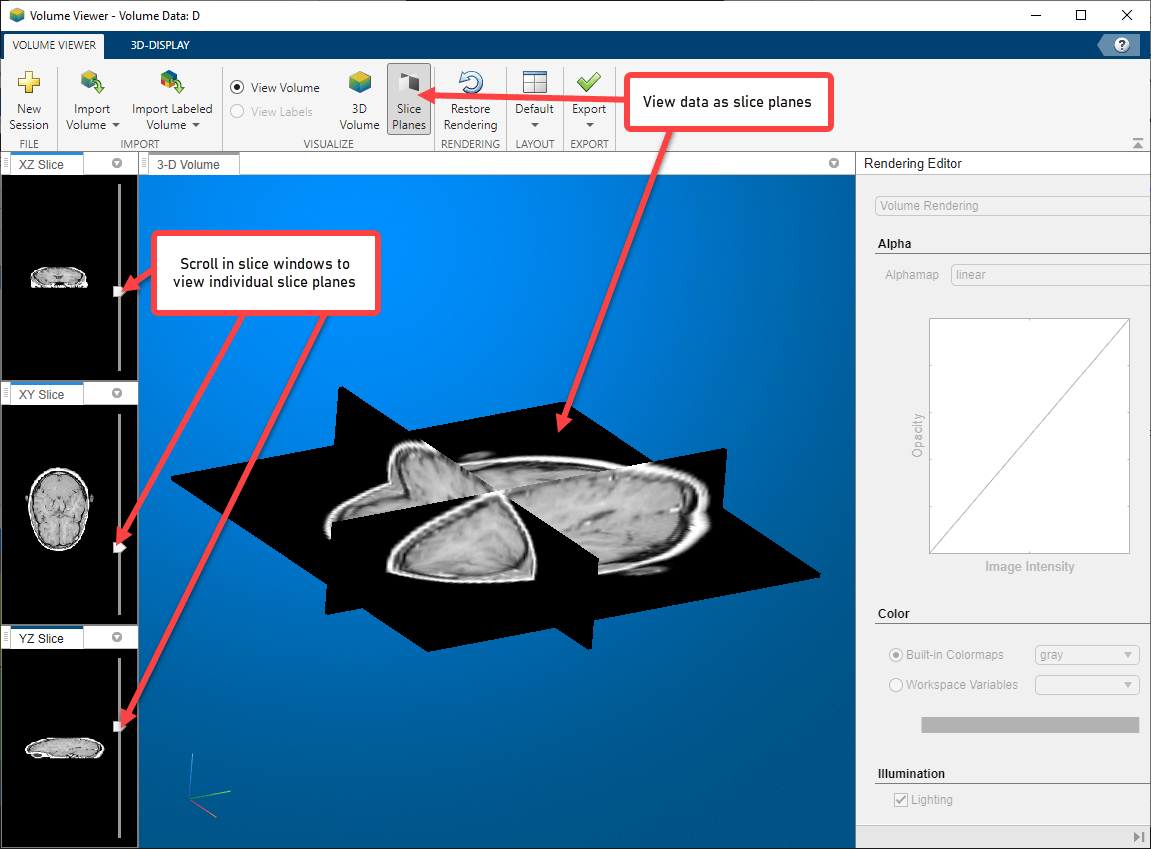
Continue using Volume Viewer capabilities until you achieve the best view of your data.
Adjust View of Volume Data in Volume Viewer
In this part of the example, you adjust the view of the volumetric data in the Volume Viewer app.
Click 3D Volume to return to viewing your data as a volume and use the capabilities of Volume Viewer to get the best visualization of your data. Volume Viewer provides several spatial referencing options that let you get a more realistic view of the head volume. (The head appears flattened in the default view.)
Specify Dimensions—In the 3D-Display tab you can specify the dimensions in the X, Y, and Z directions.

Upsample To Cube—Volume Viewer calculates a scale factor that makes the number of samples in each dimension the same as the largest dimension in the volume. This setting can make anisotropically sampled data appear scaled more correctly.
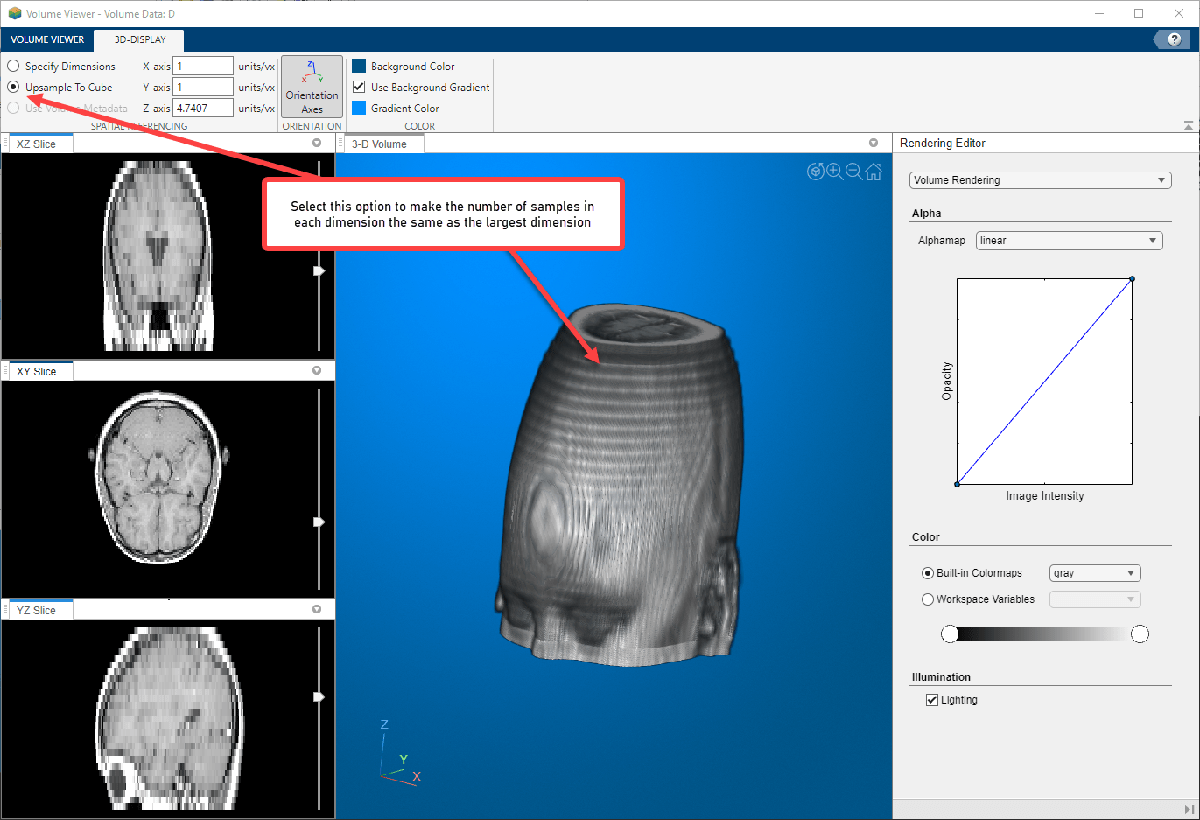
Use Volume Metadata—If the data file includes resolution data in its metadata, Volume Viewer uses the metadata and displays the volume true to scale. Volume Viewer selects the Use Volume Metadata option, by default, if metadata is present.
Refine View with Rendering Editor
This part of the example describes how to use the Volume Viewer Rendering Editor to modify your view of the data. Using the Rendering Editor, you can:
Choose the overall viewing approach: Volume Rendering, Maximum Intensity Projection, or Isosurface.
Modify the alphamap by specifying a preset alphamap, such as ct-bone, or by customizing the alphamap using the Opacity/Image Intensity curve.
Specify the colormap used in the visualization.
Specify the lighting in the visualization.
Choose the Viewing Approach
Volume Viewer offers several viewing approaches for volumes. The Maximum Intensity Projection (MIP) option looks for the voxel with the highest intensity value for each ray projected through the data. MIP can be useful for revealing the highest intensity structure within a volume. You can also view the volume as an Isosurface.
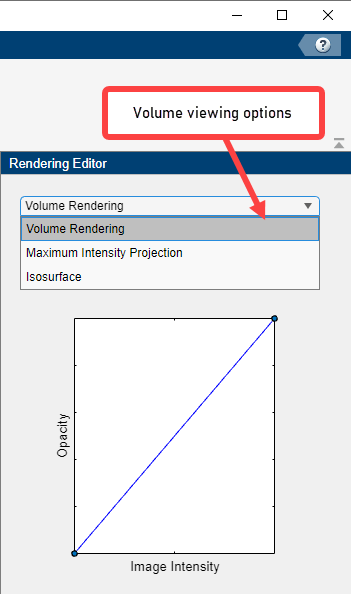
Specify the Alphamap
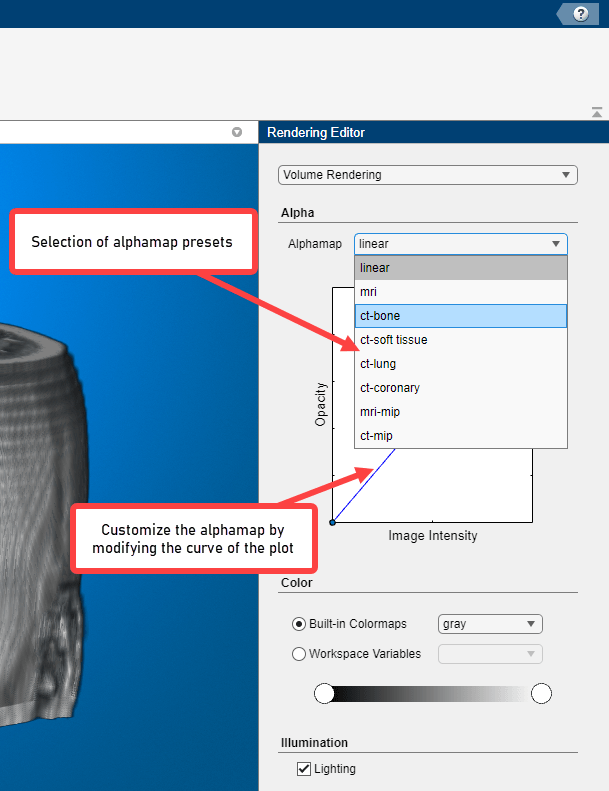
Volume rendering is highly dependent on defining an appropriate alphamap so that structures you want to see are opaque and structures you do not want to see are transparent. The Rendering Editor lets you define the opacity and transparency of voxel values throughout the volume. You can choose from a set of alphamap presets that automatically achieve certain well-defined effects. For example, to define a view that works well with CT bone data, select the CT Bone rendering preset. By default, Volume Viewer uses a simple linear relationship, but each preset changes the curve of the plot to give certain data value more or less opacity. You customize the alphamap by manipulating the plot directly.
Specify the Colormap
Color, when used with voxel intensity and opacity, is an important element of volume visualization. In the Rendering Editor, you can select from a list of predefined MATLAB colormaps, such as jet and parula. You can also specify a custom colormap that you have defined as a variable in the workspace. You can also change the colormapping for any colormap by using the interactive color bar scale. For example, to lighten the color values in a visualization, click on the color bar to create a circular slider. To modify the colormapping so that more value map to lighter colors, move the slider to the left. You can create multiple sliders on the color bar to define other colormappings.
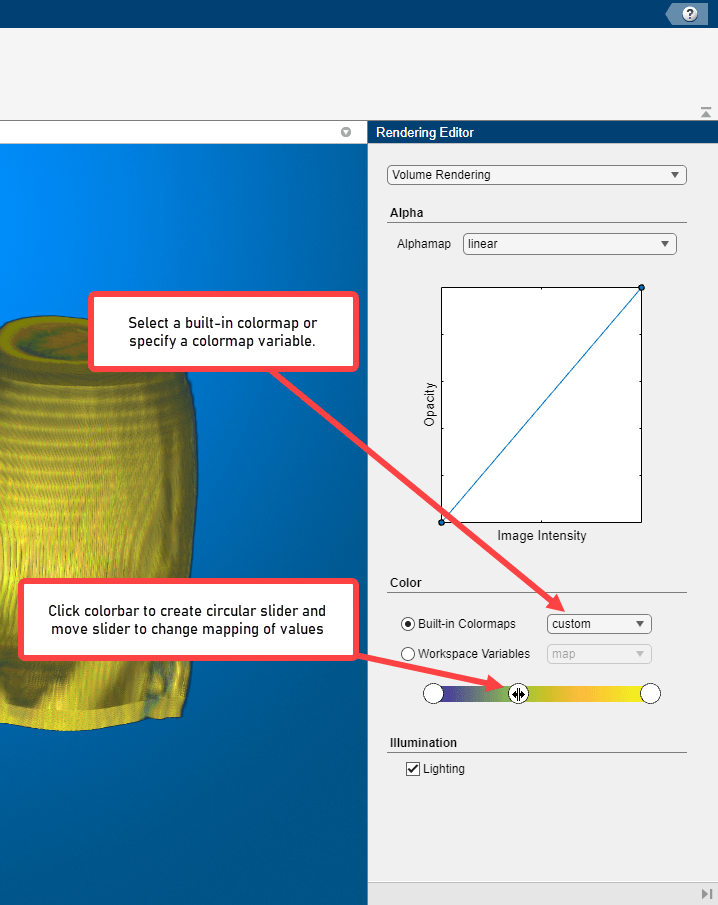
Modify Lighting Effects
By default, Volume Viewer uses certain lighting effects on the volume display. You can turn off these lighting effects by clearing the Lighting check box.
Save Volume Viewer Rendering and Camera Configuration Settings
After working in Volume Viewer to achieve the best view of your data, you can save your rendering settings and camera configuration. Volume Viewer stores this information in a structure, called config by default, that it writes to the workspace. You can use this structure with the viewer3d and volshow functions to recreate the view you achieved in Volume Viewer.
To save rendering and camera configuration settings, click Export and click the Rendering and Camera Configurations option.
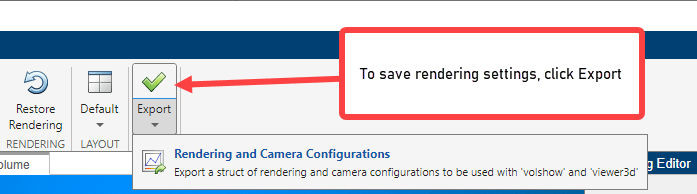
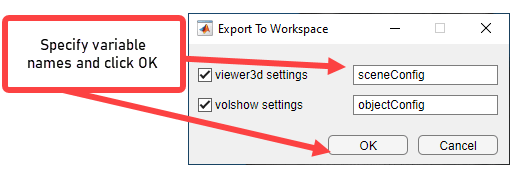
Specify the names for the structure that Volume Viewer creates or accept the default names (sceneConfig) and (objectConfig), then click OK.
References
[1] Medical Segmentation Decathlon. "Brain Tumours." Tasks. Accessed May 10, 2018. http://medicaldecathlon.com/.
The BraTS data set is provided by Medical Segmentation Decathlon under the CC-BY-SA 4.0 license. All warranties and representations are disclaimed. See the license for details. MathWorks® has modified the subset of data used in this example. This example uses the MRI data of one scan from the original data set, saved to a MAT file.
